| Step | Module | Key_files |
|---|---|---|
| 0 | Input metadata / TBI files | 0_tbi_file*.R |
| 1 | Integration by sex | NuovaPipeNo120/1_Int*_NewData.R |
| 2 | First annotation | NuovaPipeNo120/2_Ann*_New.R |
| 3 | Refined annotation | NuovaPipeNo120/3_Ann*_pt2_New.R |
| 4 | Gene filtering | NuovaPipeNo120/4_FiltGenes_*_NewData.R |
| 5 | Pseudo-bulk differential expression | NuovaPipeNo120/5_DEG*.R |
| 6 | Cell-level DE with devil | NuovaPipeNo120/6_DEA_devil_*.R |
| 7-8 | GSEA / pathway interpretation | NuovaPipeNo120/7_GSEA_F.R, 8_GSEA_* |
| 10 | Cell-cell communication | NuovaPipeNo120/10_CCC_*.R |
Cachectic Multiome Human
Interactive discussion document
Core aim
Use this page as the live presentation surface for the cachexia multiome project: start with the main biological story, then jump into QC, annotation, DEG, GSEA, DEVIL, or CCC only when someone asks.
Status
This is the scaffold. It is ready for discussion notes and exported figures; the analysis scripts remain in the repository as the reproducible backend.
Jump to what they ask
Study map QC Annotation DEG DEVIL GSEA CCC Files Open questions
Study Map
Keep each section to one answerable question, one primary visual, and one collapsible methods block. During a meeting, the document becomes a map rather than a long linear talk.
Talking path for a 5-minute overview
- What biological contrast defines cachexia in this cohort?
- Which nuclei/cell states are robust after QC and annotation?
- Which changes are shared versus sex-specific?
- Which pathways move consistently across pseudo-bulk, DEVIL, and GSEA?
- Which cell-cell communication signals are worth validating?
Quality Control
Question to answer live: are the nuclei/cells reliable enough to support the downstream story?
K017 before filtering
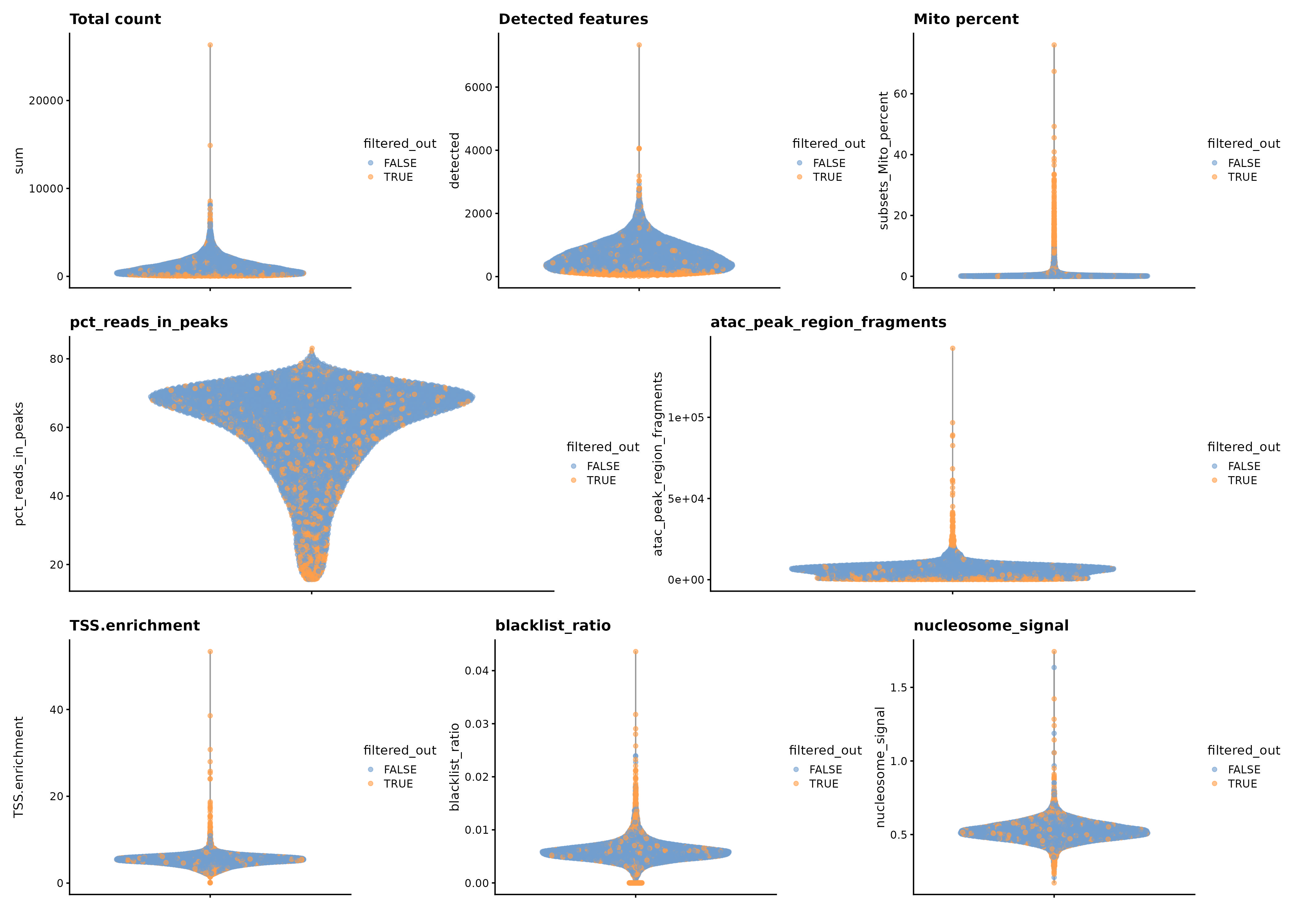
K017 after filtering
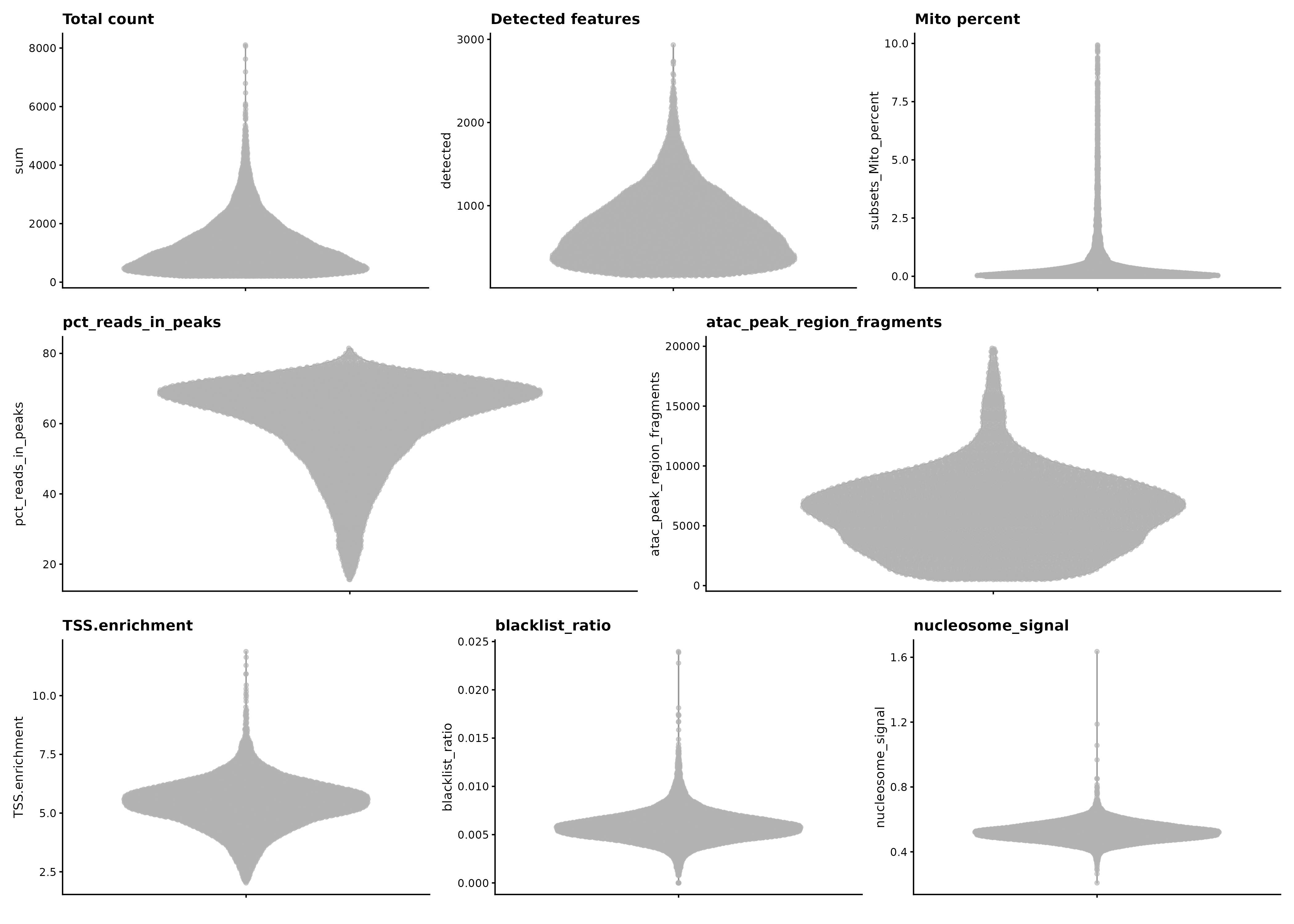
RNA QC
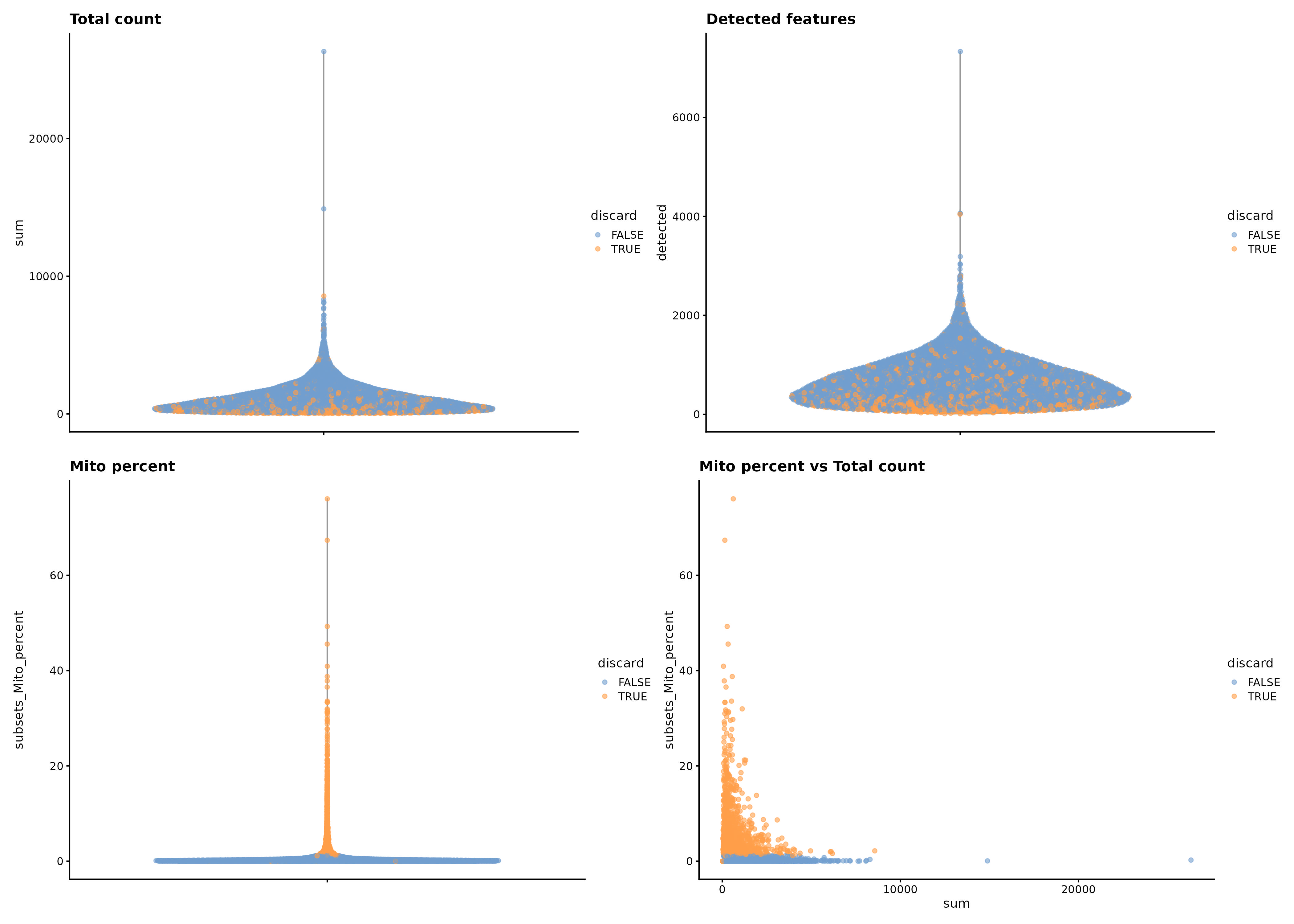
ATAC QC
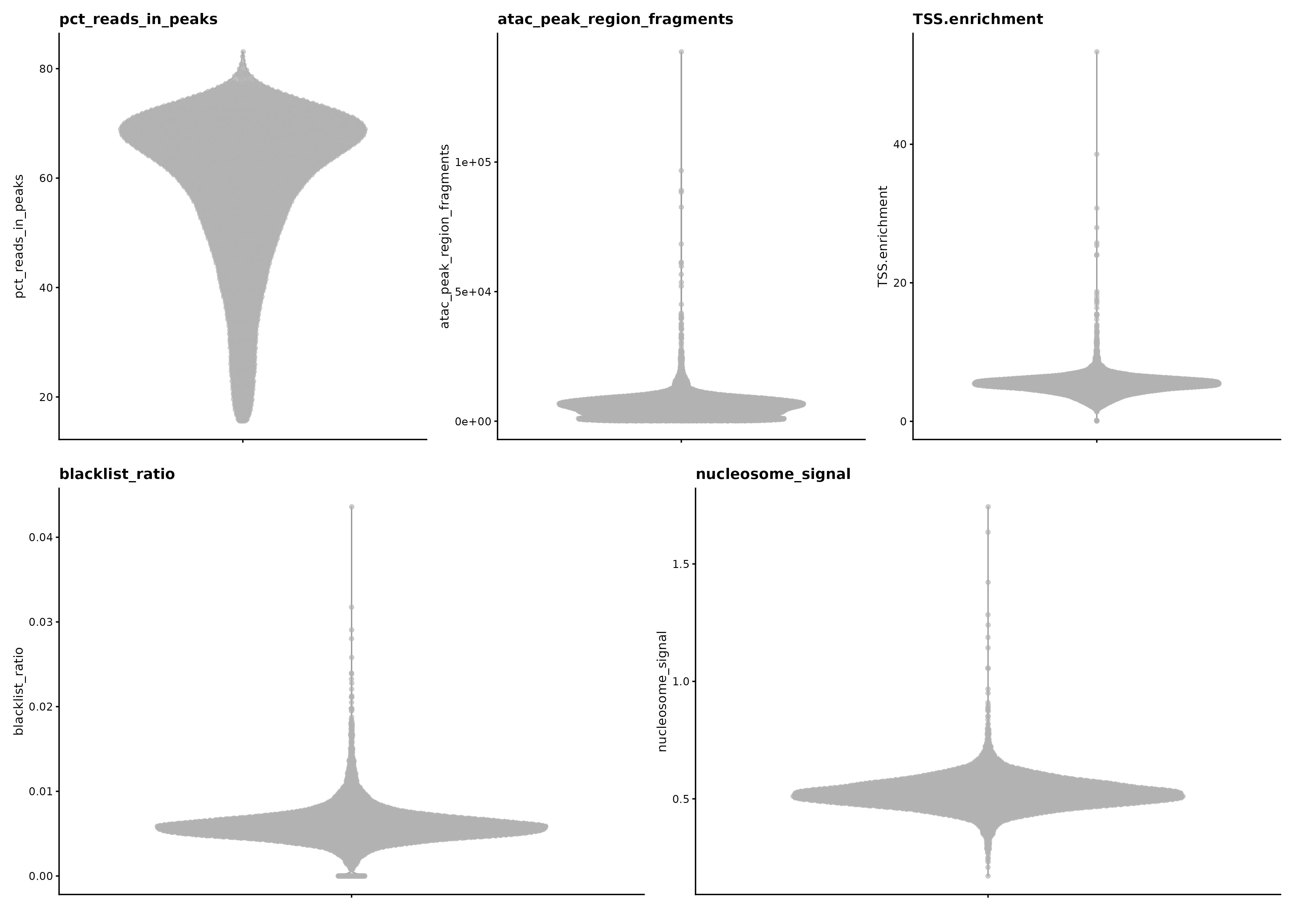
Relevant scripts and logs
| File |
|---|
| 1_QualityControl.R |
| 1_QualityControl_NewData.R |
| 2_RemoveDoublets_Fem.R |
| 2_RemoveDoublets_Fem_NewData.R |
| 2_RemoveDoublets_Man.R |
| 2_RemoveDoublets_Man_NewData.R |
| 3_QC_ribosom.R |
Annotation
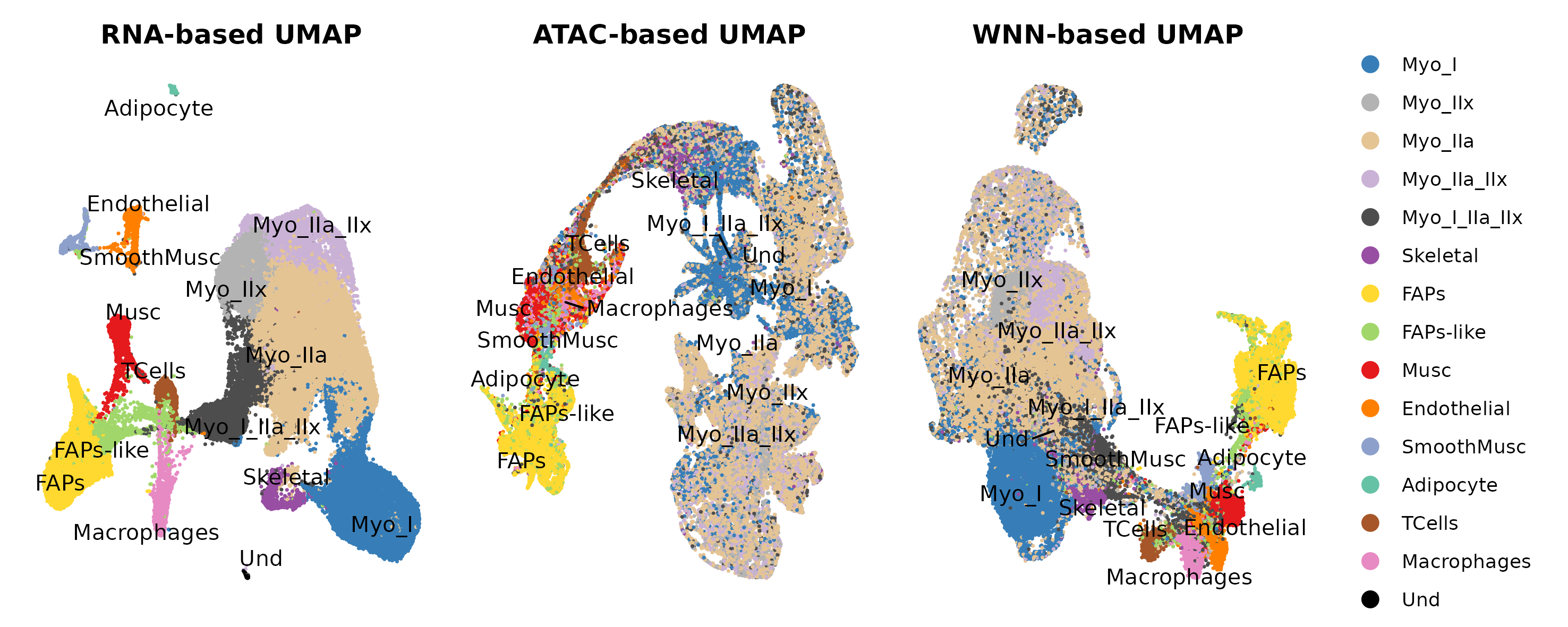
Question to answer live: what cell types/states are present, and which annotations are stable across RNA, ATAC, and WNN views?
Annotated UMAP
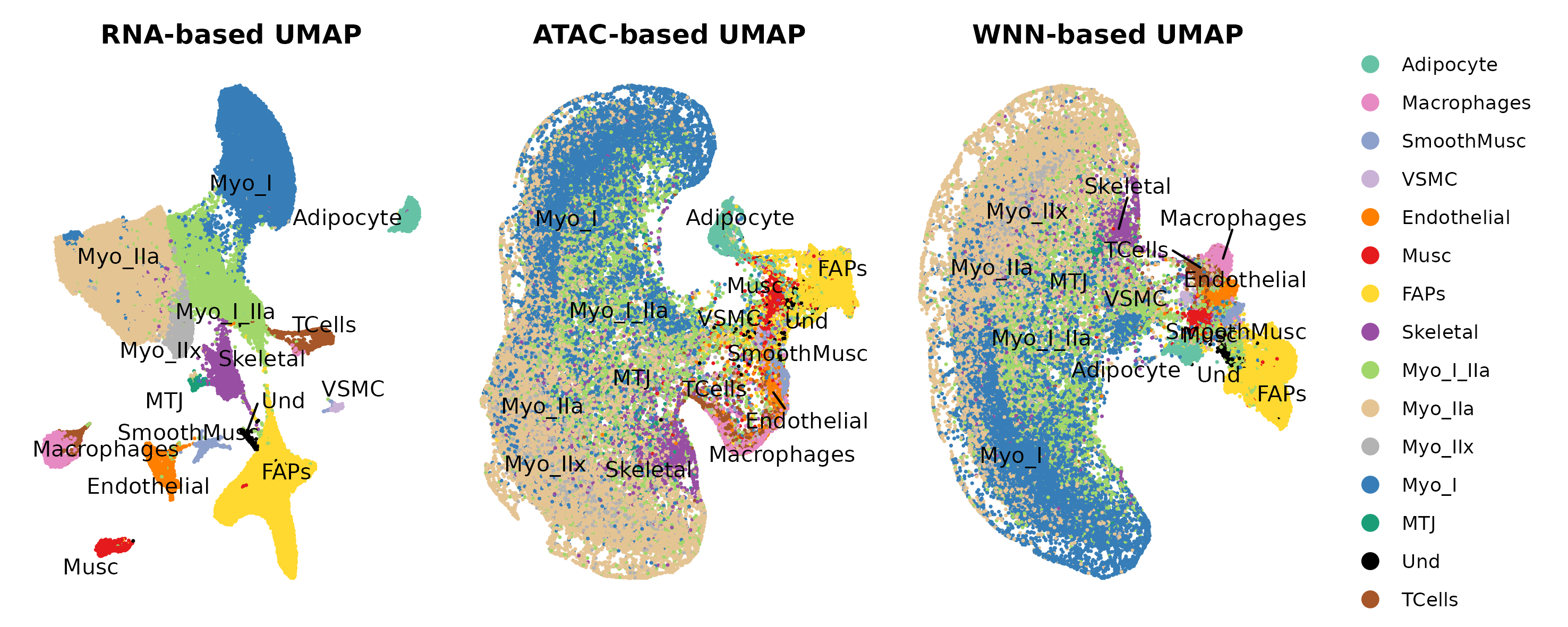
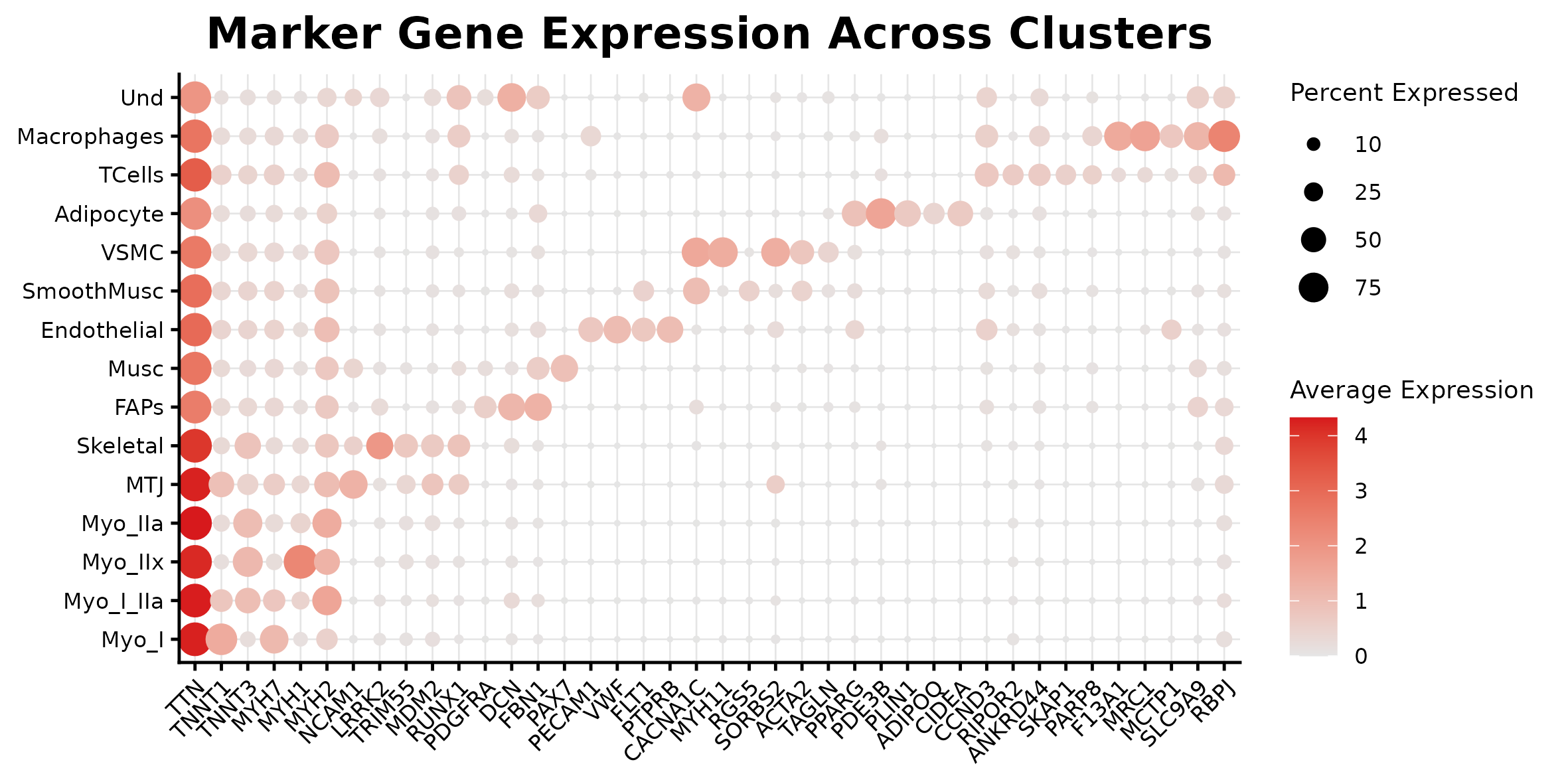
Marker dotplot
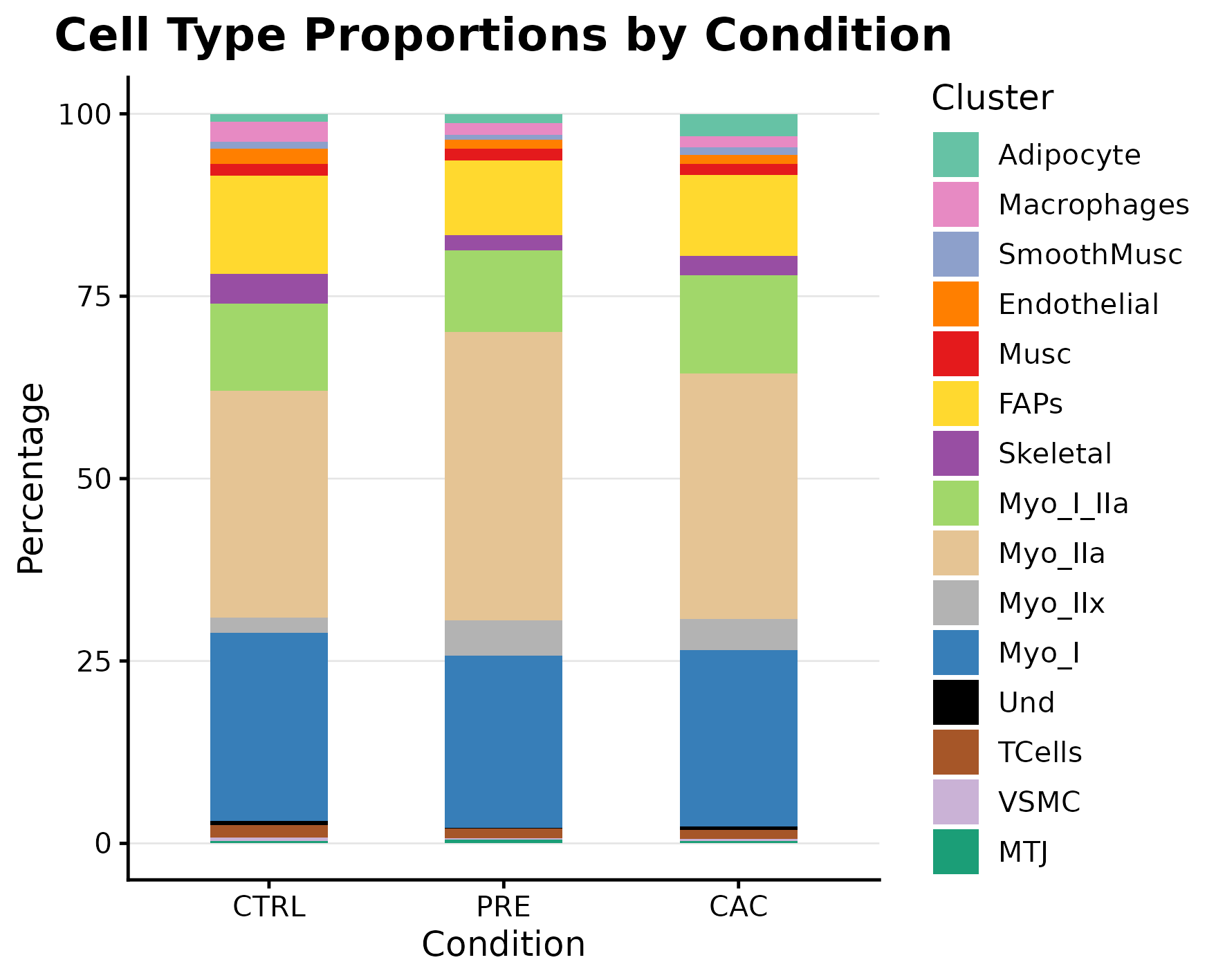
Cell composition by condition
Annotated UMAP
Marker dotplot
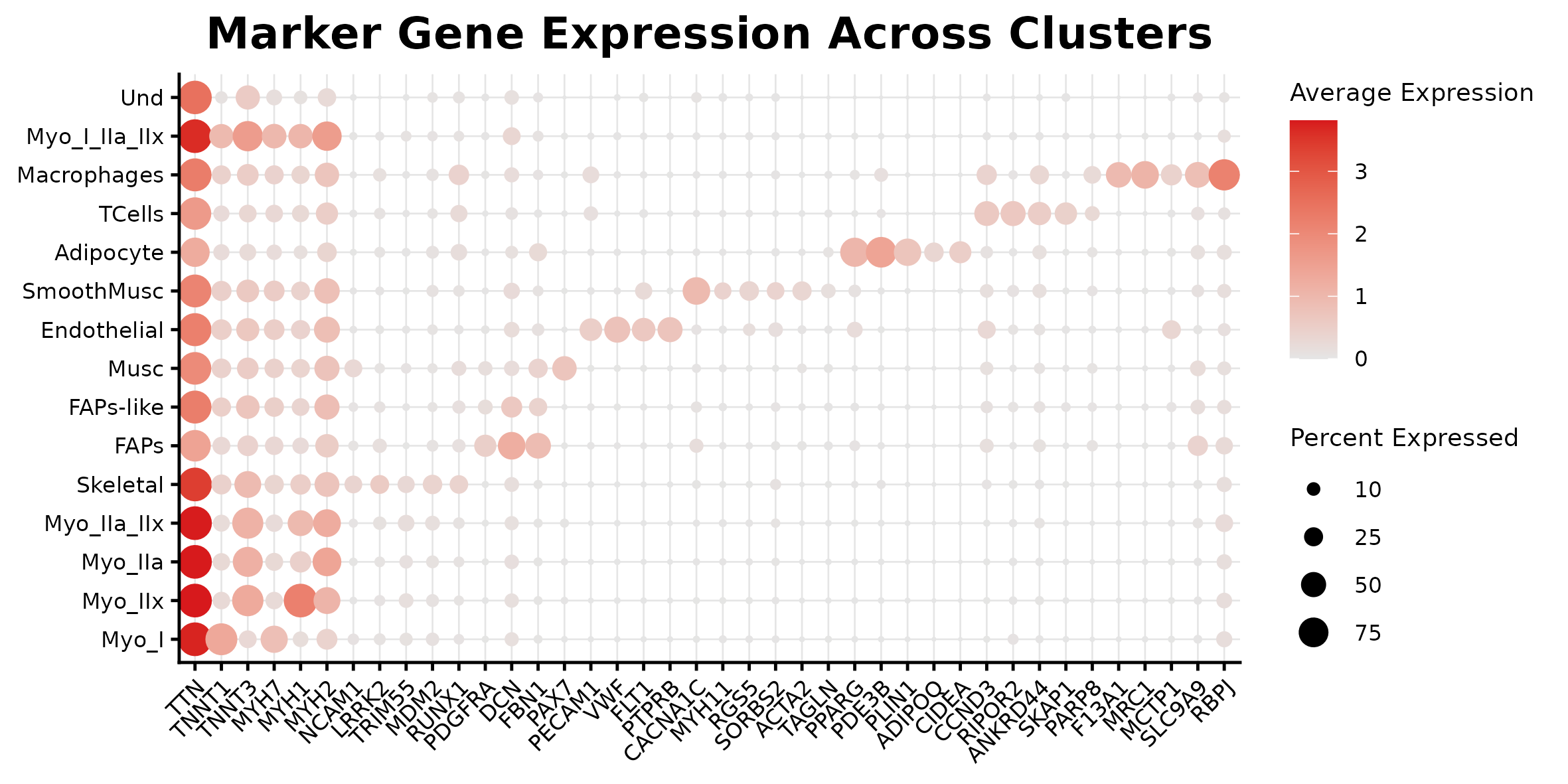
Cell composition by condition
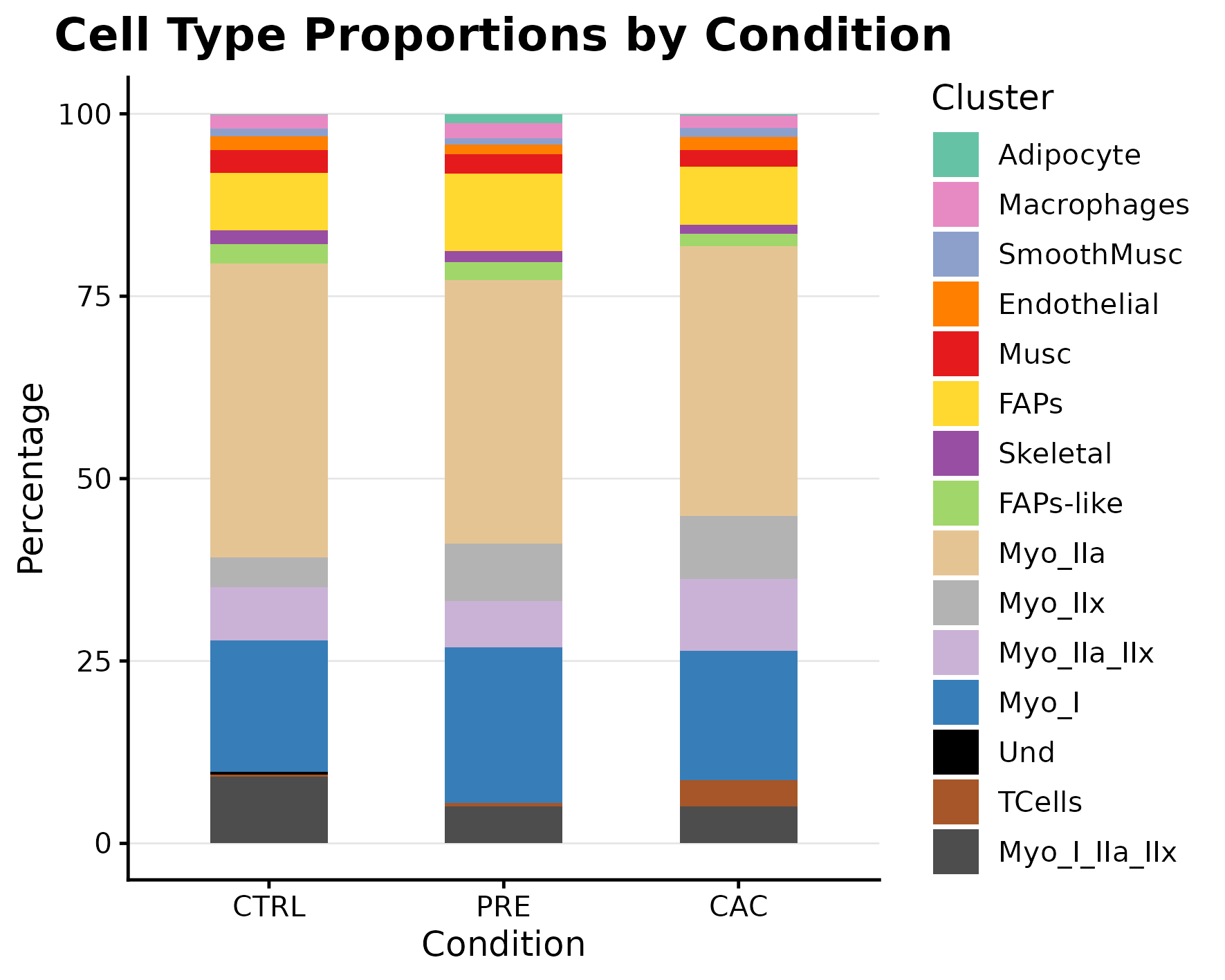
Use this tab for the curated marker list and difficult annotation decisions.
Relevant scripts
| File |
|---|
Differential Expression
Question to answer live: which transcriptional changes separate CAC, PRE, and CTRL, and are they cluster-specific or global?
The current local archive contains DEG results mainly as .xlsx tables; this tab keeps their locations close to the visual pathway summaries below.
| Table |
|---|
| F/Cluster/DEA_FEM_CAC_vs_CTRL.xlsx |
| F/Cluster/DEA_FEM_CAC_vs_PRE.xlsx |
| F/Cluster/DEA_FEM_PRE_vs_CTRL.xlsx |
| F/Clusters_NoRibo/DEA_FEM_CAC_vs_CTRL.xlsx |
| F/Clusters_NoRibo/DEA_FEM_CAC_vs_PRE.xlsx |
| F/Clusters_NoRibo/DEA_FEM_PRE_vs_CTRL.xlsx |
| F/Myo/DEA_FEM_Myo_Overall_CAC_vs_CTRL.xlsx |
| F/Myo/DEA_FEM_Myo_Overall_CAC_vs_PRE.xlsx |
| F/Myo/DEA_FEM_Myo_Overall_PRE_vs_CTRL.xlsx |
| F/Myo_NoRibo/DEA_FEM_Myo_Overall_CAC_vs_CTRL.xlsx |
| F/Myo_NoRibo/DEA_FEM_Myo_Overall_CAC_vs_PRE.xlsx |
| F/Myo_NoRibo/DEA_FEM_Myo_Overall_PRE_vs_CTRL.xlsx |
| M/Clusters/DEA_MAN_CAC_vs_CTRL.xlsx |
| M/Clusters/DEA_MAN_CAC_vs_PRE.xlsx |
| M/Clusters/DEA_MAN_PRE_vs_CTRL.xlsx |
| M/Myo/DEA_MAN_Myo_Overall_CAC_vs_CTRL.xlsx |
| M/Myo/DEA_MAN_Myo_Overall_CAC_vs_PRE.xlsx |
| M/Myo/DEA_MAN_Myo_Overall_PRE_vs_CTRL.xlsx |
| M/old/cluster/DEA_MAN_CAC_vs_CTRL.xlsx |
| M/old/cluster/DEA_MAN_CAC_vs_PRE.xlsx |
| M/old/cluster/DEA_MAN_PRE_vs_CTRL.xlsx |
| M/old/cluster_noRibo/DEA_MAN_CAC_vs_CTRL.xlsx |
| M/old/cluster_noRibo/DEA_MAN_CAC_vs_PRE.xlsx |
| M/old/cluster_noRibo/DEA_MAN_PRE_vs_CTRL.xlsx |
| M/old/Myo/DEA_MAN_Myo_Overall_CAC_vs_CTRL.xlsx |
| M/old/Myo/DEA_MAN_Myo_Overall_CAC_vs_PRE.xlsx |
| M/old/Myo/DEA_MAN_Myo_Overall_PRE_vs_CTRL.xlsx |
| M/old/Myo_NoRibo/DEA_MAN_Myo_Overall_CAC_vs_CTRL.xlsx |
| M/old/Myo_NoRibo/DEA_MAN_Myo_Overall_CAC_vs_PRE.xlsx |
| M/old/Myo_NoRibo/DEA_MAN_Myo_Overall_PRE_vs_CTRL.xlsx |
Use this tab for CAC vs CTRL, CAC vs PRE, and PRE vs CTRL in female samples.
Use this tab for CAC vs CTRL, CAC vs PRE, and PRE vs CTRL in male samples.
Use this tab for myonuclei-specific contrasts and comparisons with global/cluster-level results.
Relevant scripts and outputs
| File |
|---|
DEVIL Cell-Level DE
Question to answer live: do cell-level models support or refine the pseudo-bulk signals?
| Table |
|---|
| F/Clusters/DEA_DEVIL_FEM_CAC_vs_CTRL.xlsx |
| F/Clusters/DEA_DEVIL_FEM_CAC_vs_PRE.xlsx |
| F/Clusters/DEA_DEVIL_FEM_PRE_vs_CTRL.xlsx |
| F/Clusters_NoRibo/DEA_DEVIL_FEM_CAC_vs_CTRL.xlsx |
| F/Clusters_NoRibo/DEA_DEVIL_FEM_CAC_vs_PRE.xlsx |
| F/Clusters_NoRibo/DEA_DEVIL_FEM_PRE_vs_CTRL.xlsx |
| F/Custom_Sample_Comparisons/DEA_DEVIL_FEM_CUSTOM_All_CTRL_vs_K048.xlsx |
| F/Custom_Sample_Comparisons/DEA_DEVIL_FEM_CUSTOM_All_CTRL_vs_PK120.xlsx |
| F/Myo/DEA_DEVIL_FEM_Myo_Overall_CAC_vs_CTRL.xlsx |
| F/Myo/DEA_DEVIL_FEM_Myo_Overall_CAC_vs_PRE.xlsx |
| F/Myo/DEA_DEVIL_FEM_Myo_Overall_PRE_vs_CTRL.xlsx |
| F/Myo_NoRibo/DEA_DEVIL_FEM_Myo_Overall_CAC_vs_CTRL.xlsx |
| F/Myo_NoRibo/DEA_DEVIL_FEM_Myo_Overall_CAC_vs_PRE.xlsx |
| F/Myo_NoRibo/DEA_DEVIL_FEM_Myo_Overall_PRE_vs_CTRL.xlsx |
| M/Clusters/DEA_DEVIL_MAN_CAC_vs_CTRL.xlsx |
| M/Clusters/DEA_DEVIL_MAN_CAC_vs_PRE.xlsx |
| M/Clusters/DEA_DEVIL_MAN_PRE_vs_CTRL.xlsx |
| M/Myo/DEA_DEVIL_MAN_Myo_Overall_CAC_vs_CTRL.xlsx |
| M/Myo/DEA_DEVIL_MAN_Myo_Overall_CAC_vs_PRE.xlsx |
| M/Myo/DEA_DEVIL_MAN_Myo_Overall_PRE_vs_CTRL.xlsx |
The visual DEVIL-ranked pathway summaries are shown under GSEA and pathways, in the DEVIL-ranked GSEA tab.
Relevant scripts and logs
| File |
|---|
GSEA And Pathways
Question to answer live: which pathways make the gene-level changes biologically coherent?
Myonuclei CAC vs CTRL
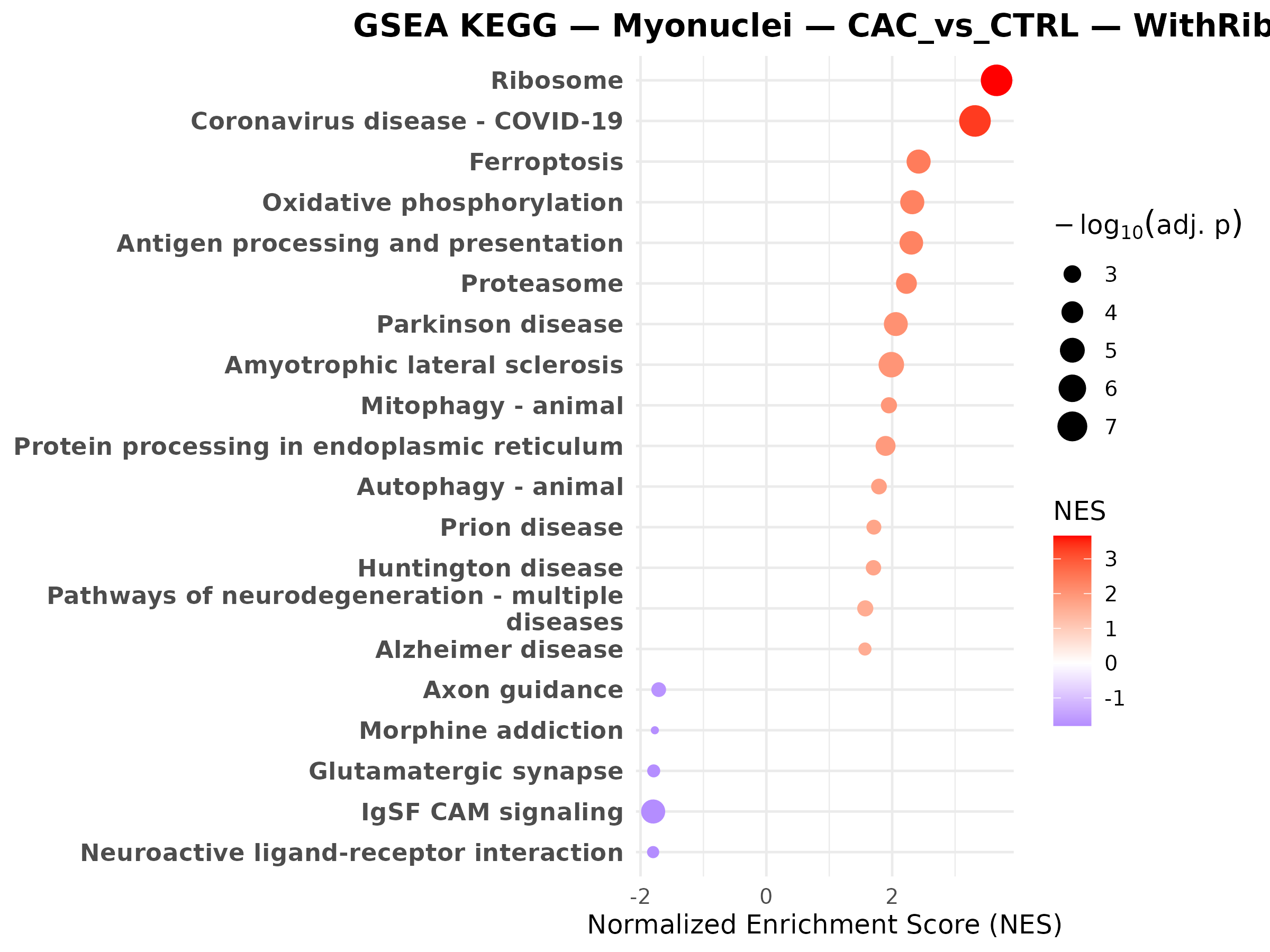
Myonuclei CAC vs PRE
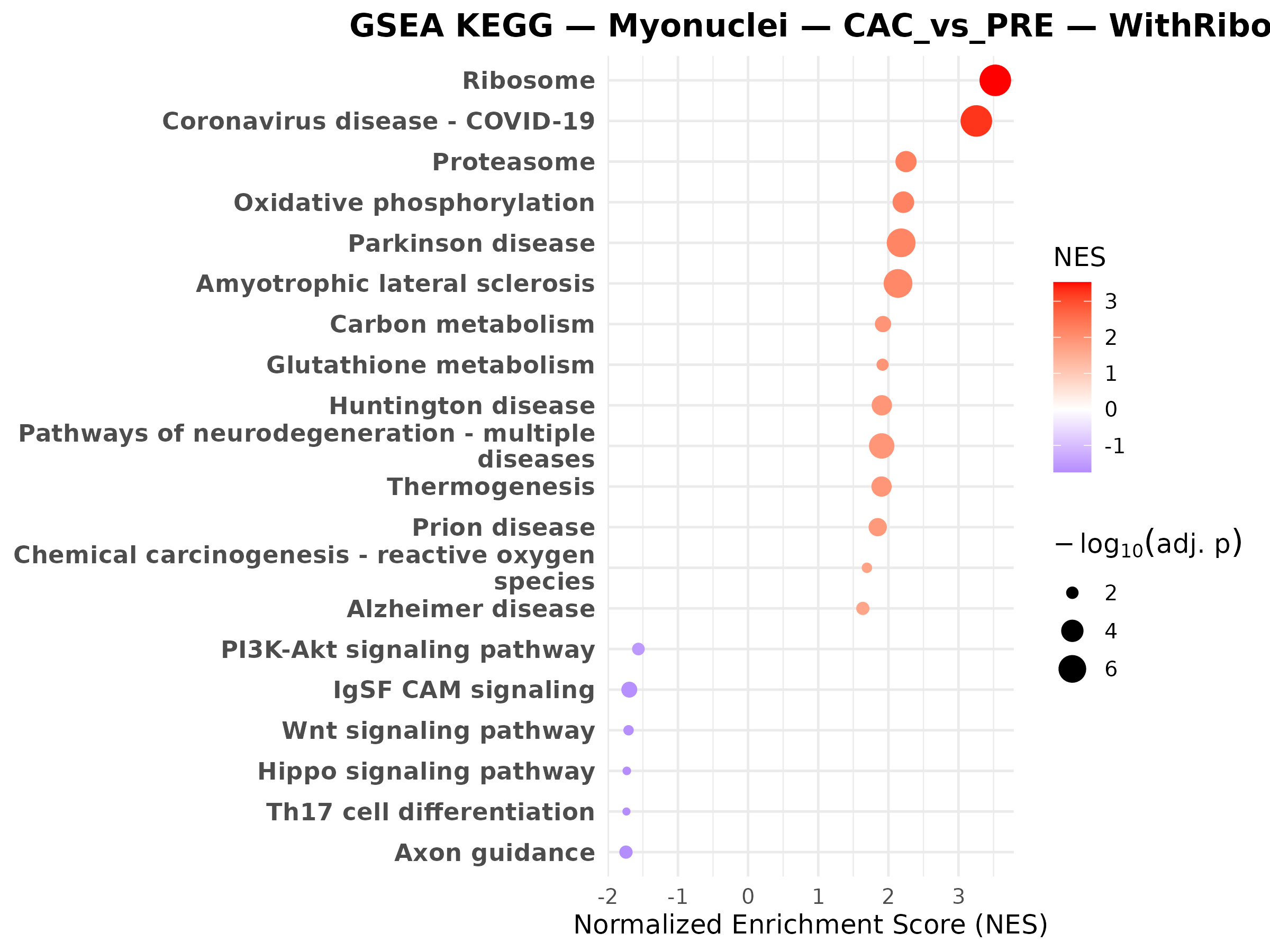
Myonuclei PRE vs CTRL
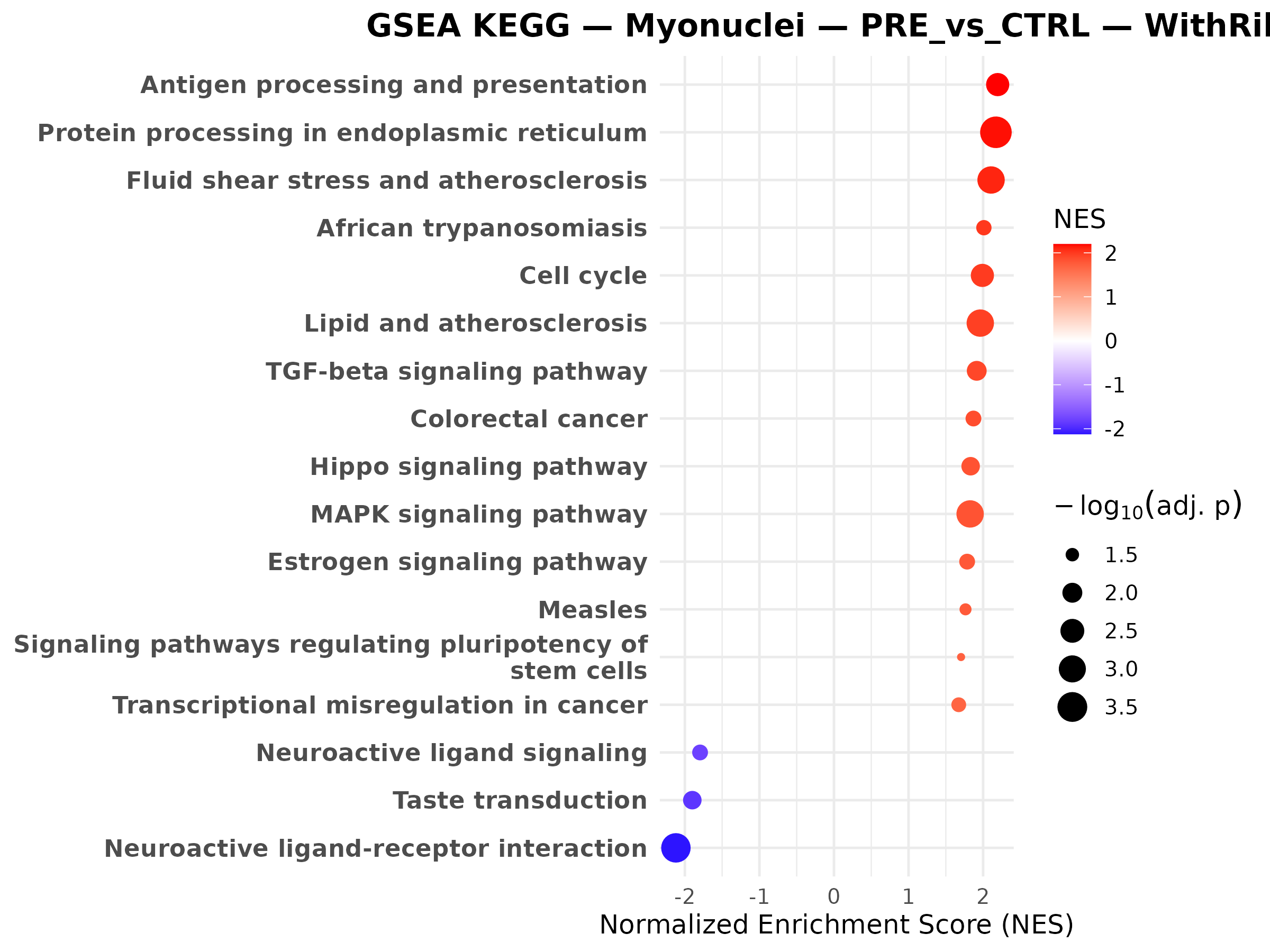
Myonuclei CAC vs CTRL
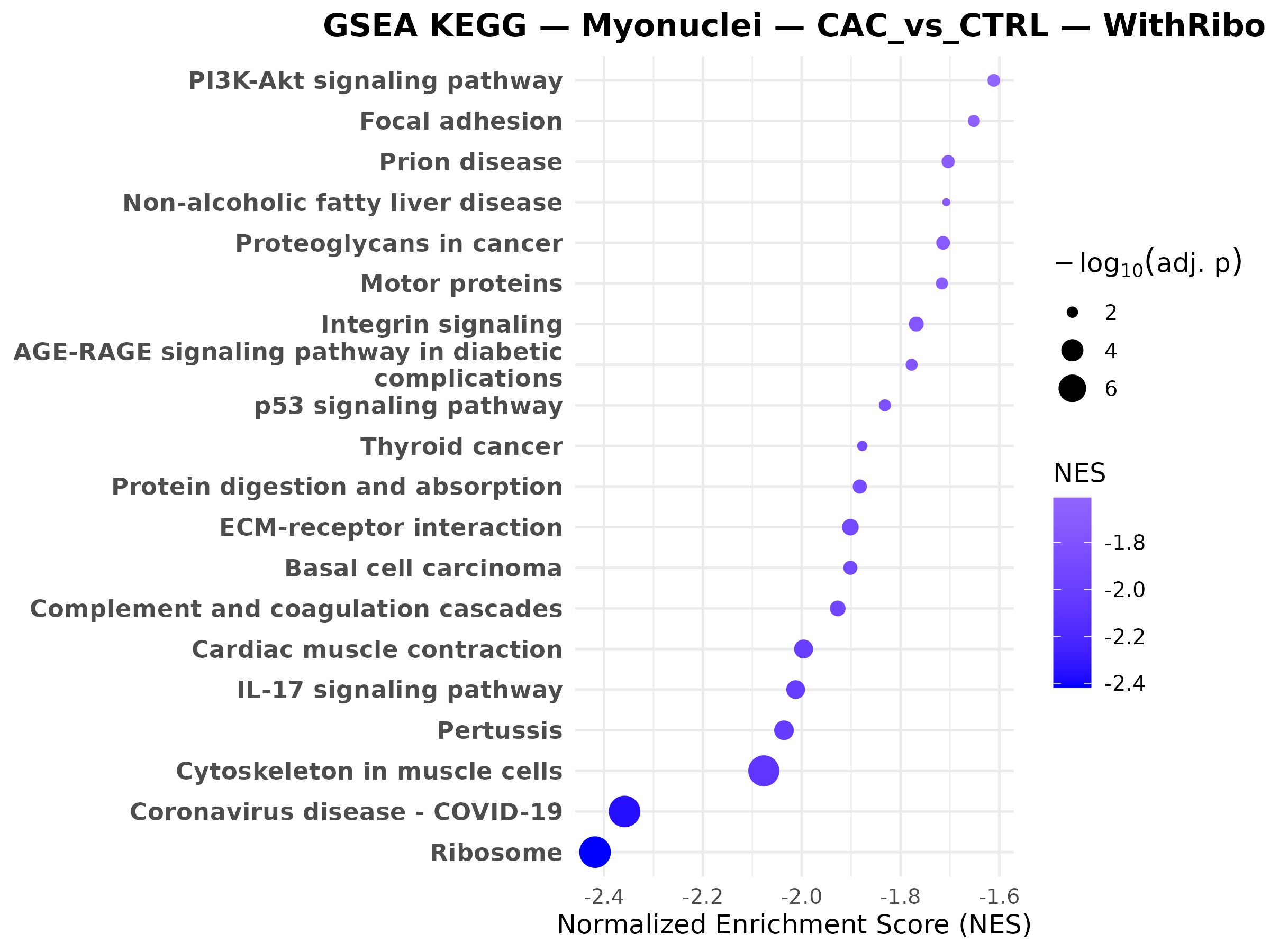
Myonuclei CAC vs PRE
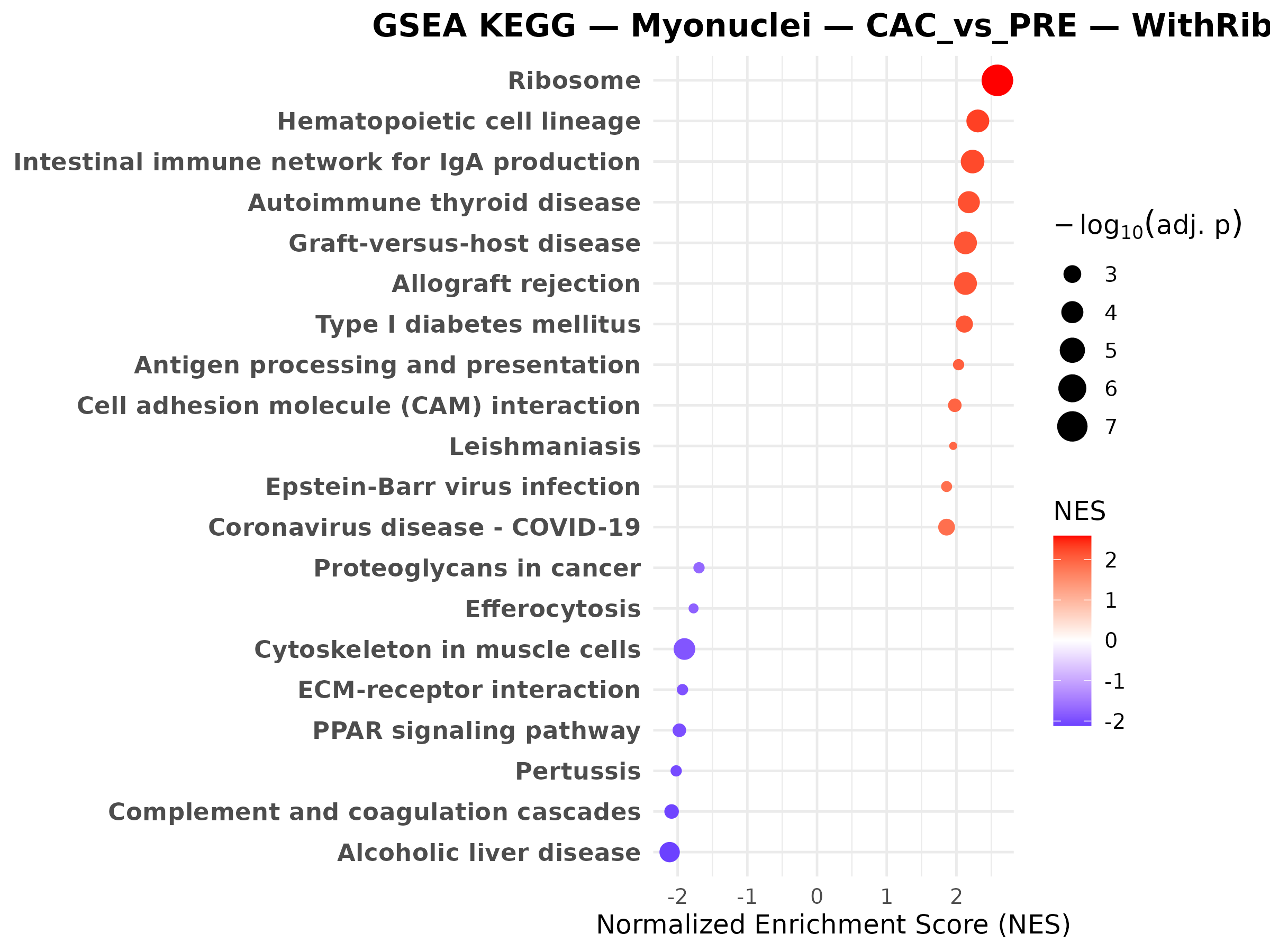
Myonuclei PRE vs CTRL
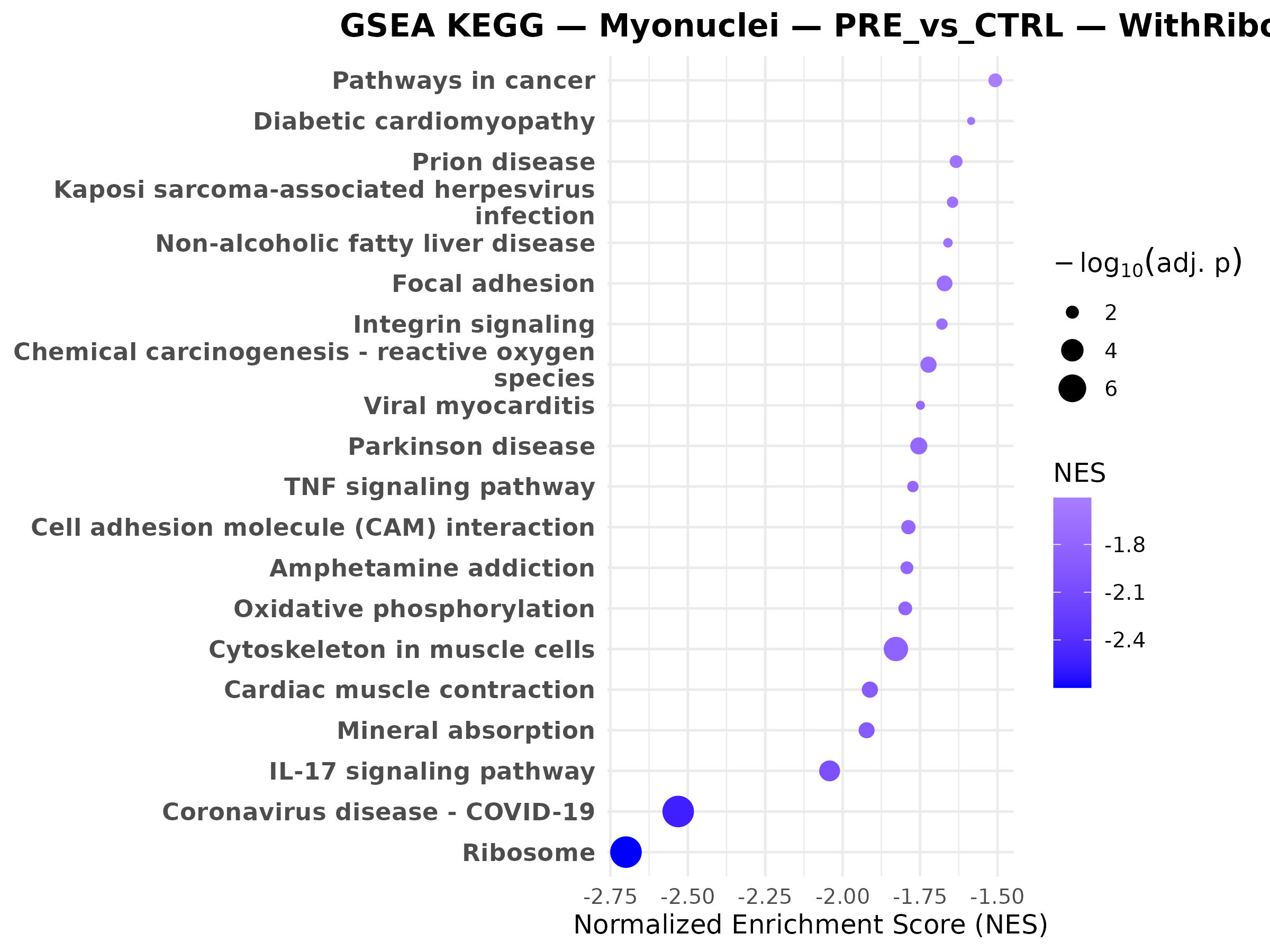
Female myonuclei CAC vs CTRL
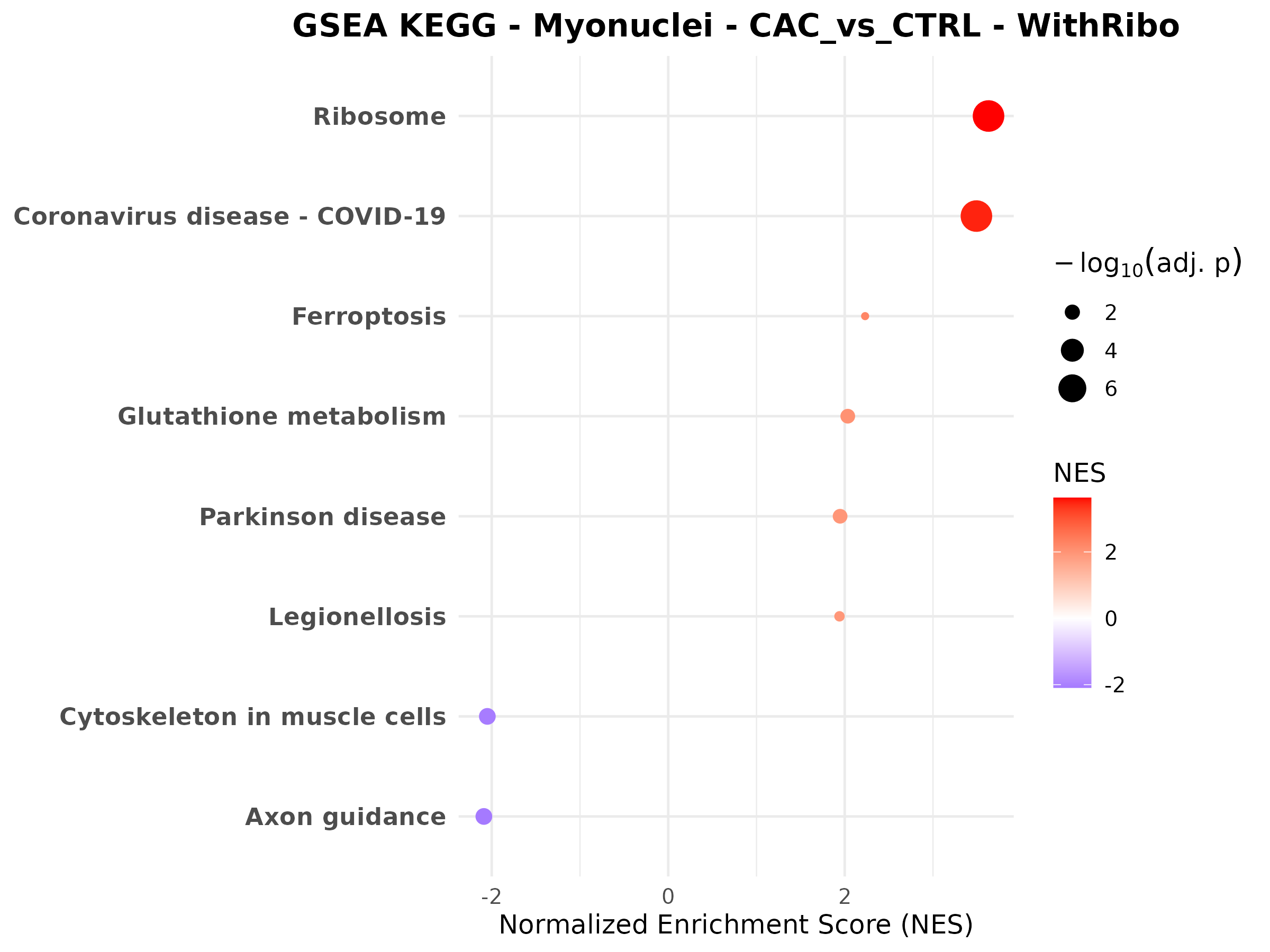
Male myonuclei CAC vs CTRL
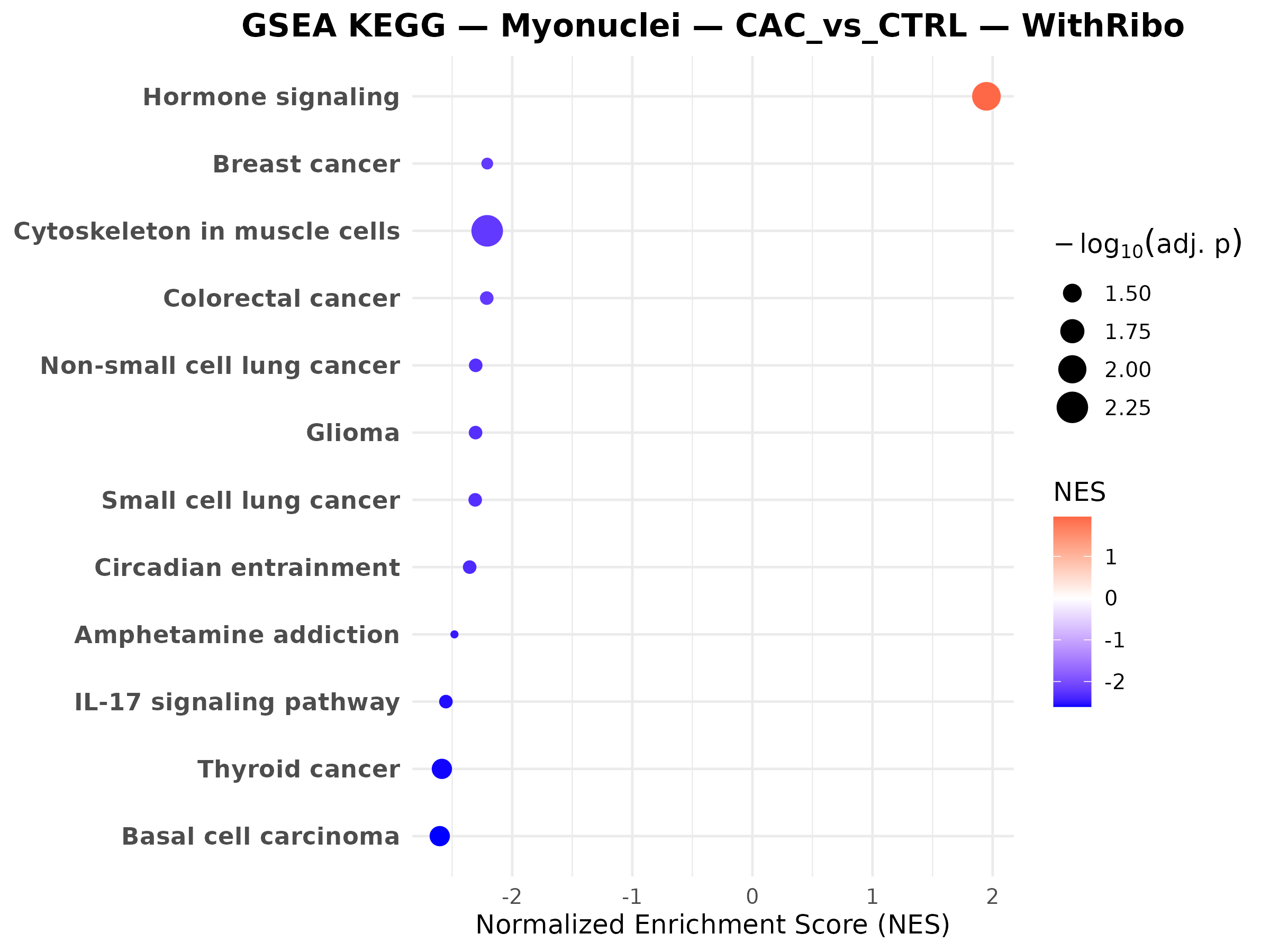
Relevant scripts and logs
| File |
|---|
Cell-Cell Communication
Question to answer live: which ligand-receptor programs connect cachexia-associated states?
CAC Myo IIx to T cells: clusters
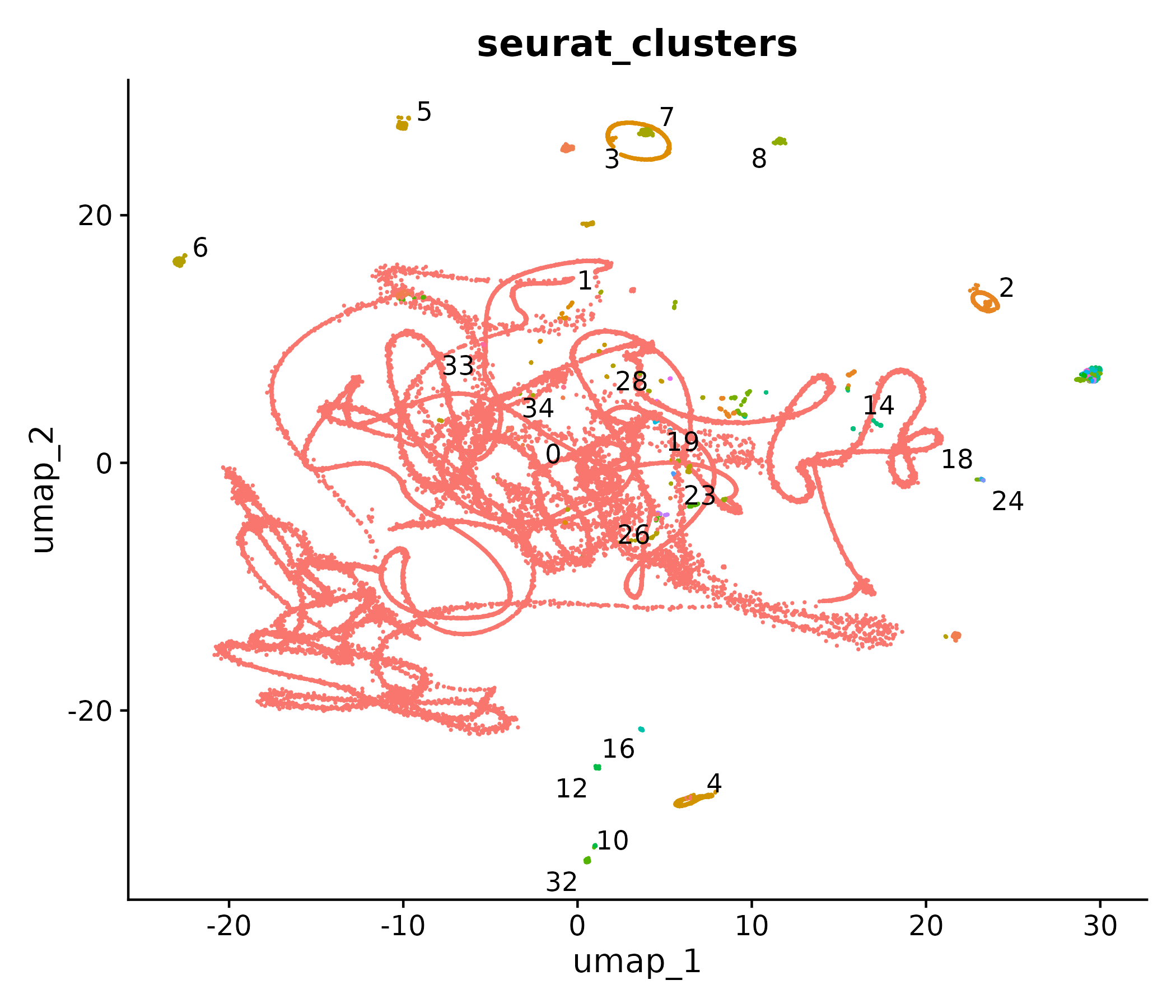
Sender clusters
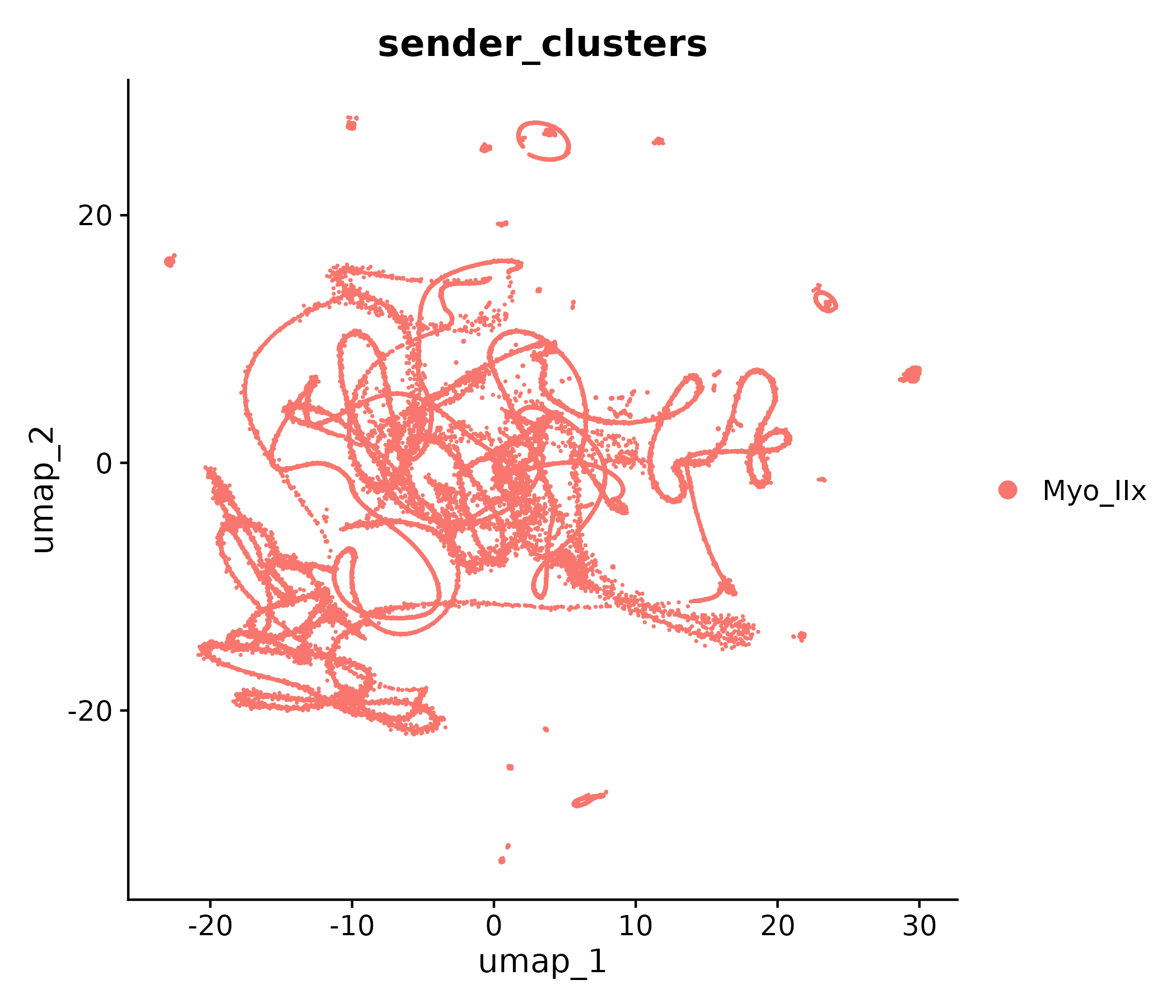
Receiver clusters
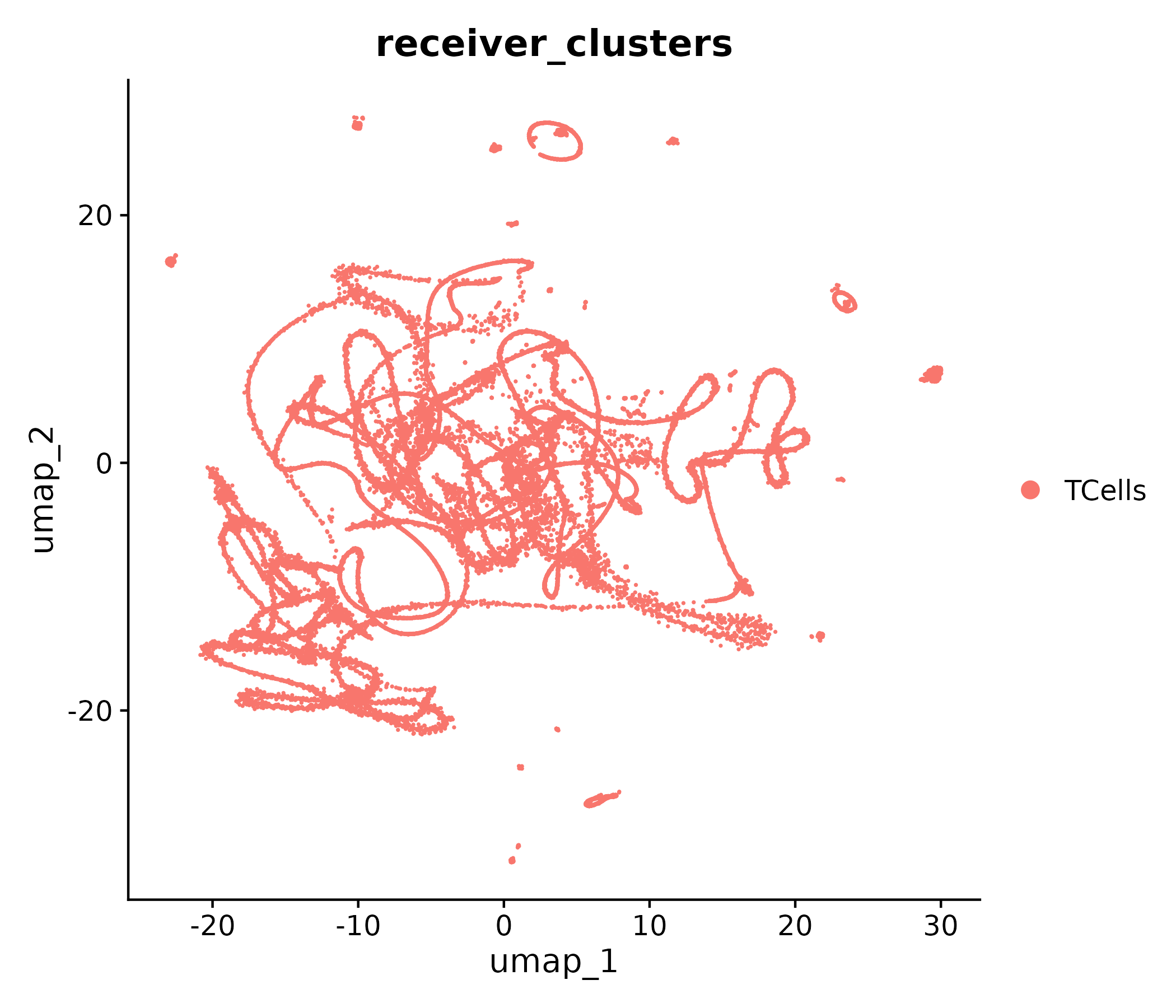
Top ligand-receptor markers
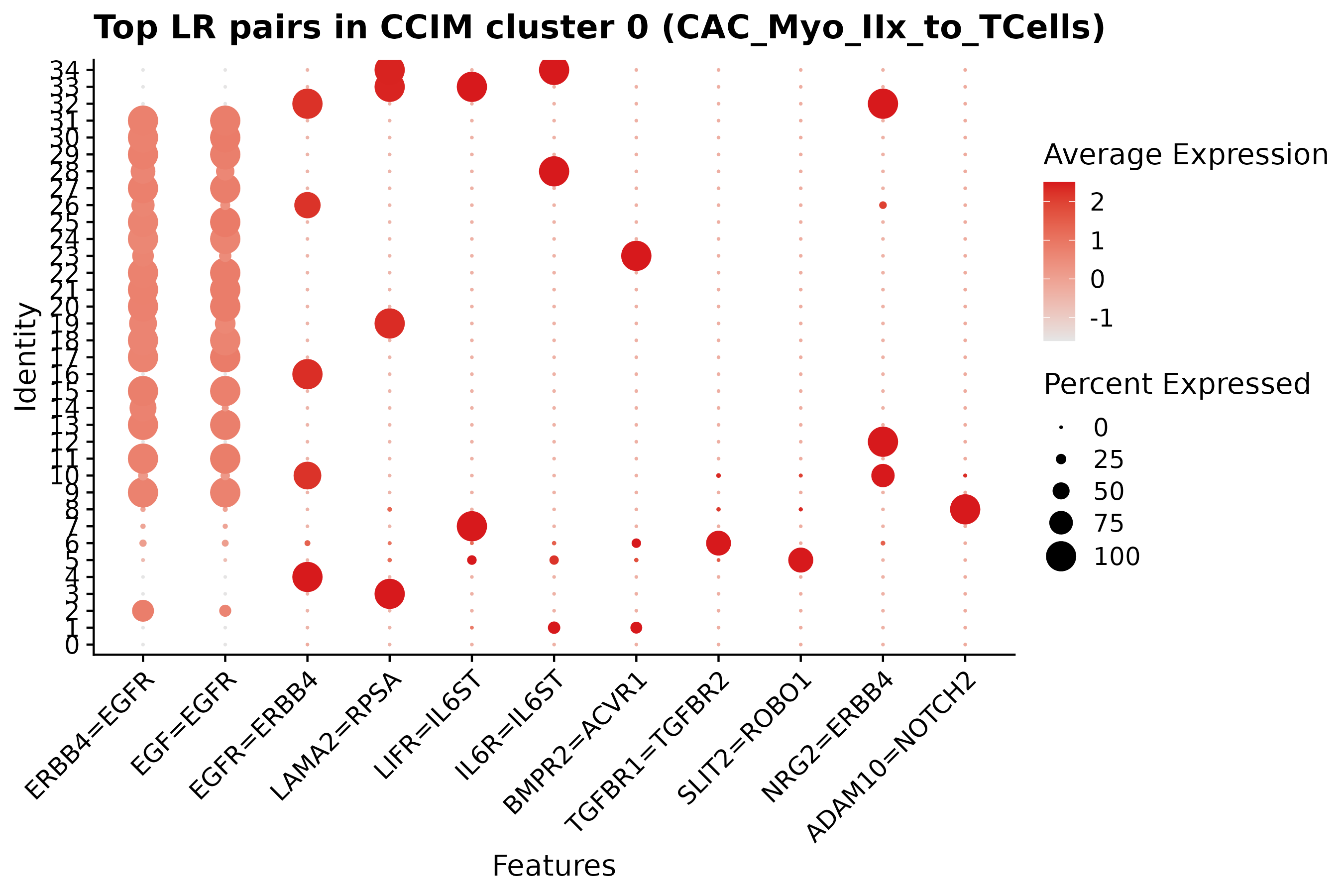
Relevant scripts and logs
| File |
|---|
Figure Gallery
### annotation_f_cells_by_condition.png
{fig-alt='annotation_f_cells_by_condition.png'}
### annotation_f_marker_dotplot.png
{fig-alt='annotation_f_marker_dotplot.png'}
### annotation_f_umap_combined.png
{fig-alt='annotation_f_umap_combined.png'}
### annotation_m_cells_by_condition.png
{fig-alt='annotation_m_cells_by_condition.png'}
### annotation_m_marker_dotplot.png
{fig-alt='annotation_m_marker_dotplot.png'}
### annotation_m_umap_combined.png
{fig-alt='annotation_m_umap_combined.png'}
### ccc_myoIIx_to_tcells_clusters.png
{fig-alt='ccc_myoIIx_to_tcells_clusters.png'}
### ccc_myoIIx_to_tcells_lr_dotplot.png
{fig-alt='ccc_myoIIx_to_tcells_lr_dotplot.png'}
### ccc_myoIIx_to_tcells_receiver.png
{fig-alt='ccc_myoIIx_to_tcells_receiver.png'}
### ccc_myoIIx_to_tcells_sender.png
{fig-alt='ccc_myoIIx_to_tcells_sender.png'}
### devil_gsea_f_myonuclei_cac_vs_ctrl.png
{fig-alt='devil_gsea_f_myonuclei_cac_vs_ctrl.png'}
### devil_gsea_m_myonuclei_cac_vs_ctrl.png
{fig-alt='devil_gsea_m_myonuclei_cac_vs_ctrl.png'}
### gsea_f_myonuclei_cac_vs_ctrl.png
{fig-alt='gsea_f_myonuclei_cac_vs_ctrl.png'}
### gsea_f_myonuclei_cac_vs_pre.png
{fig-alt='gsea_f_myonuclei_cac_vs_pre.png'}
### gsea_f_myonuclei_pre_vs_ctrl.png
{fig-alt='gsea_f_myonuclei_pre_vs_ctrl.png'}
### gsea_m_myonuclei_cac_vs_ctrl.png
{fig-alt='gsea_m_myonuclei_cac_vs_ctrl.png'}
### gsea_m_myonuclei_cac_vs_pre.png
{fig-alt='gsea_m_myonuclei_cac_vs_pre.png'}
### gsea_m_myonuclei_pre_vs_ctrl.png
{fig-alt='gsea_m_myonuclei_pre_vs_ctrl.png'}
### qc_K017_atac.jpg
{fig-alt='qc_K017_atac.jpg'}
### qc_K017_filtered.jpg
{fig-alt='qc_K017_filtered.jpg'}
### qc_K017_rna.jpg
{fig-alt='qc_K017_rna.jpg'}
### qc_K017_unfiltered.jpg
{fig-alt='qc_K017_unfiltered.jpg'}Legacy Presentations
The old slide decks stay outside the Git clone for now. This table indexes the local presentation archive so the useful figures can be migrated gradually into this Quarto document.
| File | Type | Size_MB | Modified | |
|---|---|---|---|---|
| 46 | 14_Presentazione_13_04/Presentazione_02-03-26.pptx | PPTX | 40.7 | 2026-04-13 |
| 45 | 13_Presentrazione_23-03/Presentazione_02-03-26.pptx | PPTX | 9.1 | 2026-03-22 |
| 44 | 12_Presentazione_02-03-26/Presentazione_02-03-26.pptx | PPTX | 15.9 | 2026-03-03 |
| 43 | 12_Presentazione_02-03-26/Presentazione_02-03-26.pdf | 7.6 | 2026-03-02 | |
| 58 | 5-Presentazione-02-10-25/5-Presentazione_02-10-2025.pptx | PPTX | 21.1 | 2026-02-26 |
| 42 | 11_Presentazione_11-02-26/Presentazione_11-02-26.pptx | PPTX | 10.9 | 2026-02-23 |
| 41 | 11_Presentazione_11-02-26/Presentazione_11-02-26.pdf | 6.1 | 2026-02-11 | |
| 70 | Annotazione.pptx | PPTX | 2.8 | 2026-02-09 |
| 40 | 10_Presentazione_22-01-26/Presentazione_22-01-26.pptx | PPTX | 3.5 | 2026-01-21 |
| 66 | 9_Presentazione_Francesi-08-01-26/08-01.pptx | PPTX | 11.0 | 2026-01-08 |
| 69 | 9_Presentazione_Francesi-08-01-26/Summary.pdf | 0.1 | 2026-01-07 | |
| 68 | 9_Presentazione_Francesi-08-01-26/Script_Presentazione.pdf | 0.1 | 2026-01-07 | |
| 67 | 9_Presentazione_Francesi-08-01-26/QC.pdf | 0.1 | 2026-01-07 | |
| 65 | 8_Presentazione_18-12-25/8-Presentazione_18-12-2025.pptx | PPTX | 5.1 | 2025-12-17 |
| 64 | 7_Presentazione_04-12-25/7-Presentazione_04-12-2025.pptx | PPTX | 58.1 | 2025-12-04 |
| 63 | 7_Presentazione_04-12-25/7-CellChat_04-12-2025.pptx | PPTX | 32.2 | 2025-12-01 |
| 84 | Extra/6-Stabilita_Cluster.pdf | 3.0 | 2025-11-06 | |
| 61 | 6-Presentazione-06-11-25/6-Presentazione_OLD_06-11-2025.pptx | PPTX | 107.1 | 2025-11-06 |
| 60 | 6-Presentazione-06-11-25/6-Presentazione_06-11-2025.pptx | PPTX | 63.0 | 2025-11-06 |
| 59 | 6-Presentazione-06-11-25/6-Presentazione_06-11-2025.pdf | 34.9 | 2025-11-06 | |
| 85 | Extra/6-Stabilita_Cluster.pptx | PPTX | 4.2 | 2025-10-27 |
| 62 | 6-Presentazione-06-11-25/6-Scriabin.pptx | PPTX | 2.1 | 2025-10-27 |
| 57 | 5-Presentazione-02-10-25/5-Presentazione_02-10-2025.pdf | 14.7 | 2025-10-02 | |
| 82 | Extra/2_01-08/GSEA_Male_ComBat/2_GSEA_Male_31-07.pptx | PPTX | 3.9 | 2025-08-02 |
| 81 | Extra/2_01-08/GSEA_Male_ComBat/2_GSEA_Male.pdf | 3.3 | 2025-08-02 | |
| 79 | Extra/2_01-08/GSEA_Female_RUV/No120/2_GSEA_Female_No120.pdf | 4.2 | 2025-08-02 | |
| 80 | Extra/2_01-08/GSEA_Female_RUV/No120/2_GSEA_No120_31-07.pptx | PPTX | 5.0 | 2025-08-01 |
| 77 | Extra/2_01-08/GSEA_Female_RUV/Best/2_GSEA_Female_No120_No63.pdf | 4.0 | 2025-08-01 | |
| 74 | Extra/2_01-08/1_DEGs_IndagineCombinazioni_Summary.pdf | 0.8 | 2025-08-01 | |
| 73 | Extra/2_01-08/1_DEGs_IndagineCombinazioni.pdf | 8.4 | 2025-08-01 |
Copied asset manifest
| Section | Asset | Source |
|---|---|---|
| QC | qc/qc_K017_unfiltered.jpg | 0_QC_graph_new/S65235_K017_QC_unfiltered.jpg |
| QC | qc/qc_K017_filtered.jpg | 0_QC_graph_new/S65235_K017_QC_filtered.jpg |
| QC | qc/qc_K017_rna.jpg | 0_QC_graph_new/S65235_K017_QC_RNA.jpg |
| QC | qc/qc_K017_atac.jpg | 0_QC_graph_new/S65235_K017_QC_ATAC.jpg |
| Annotation | annotation/annotation_f_umap_combined.png | 3_Ann_pt2/F/UMAP_annot_combined.png |
| Annotation | annotation/annotation_f_marker_dotplot.png | 3_Ann_pt2/F/Multiome_DotPlot_Markers.png |
| Annotation | annotation/annotation_f_cells_by_condition.png | 3_Ann_pt2/F/BarPlot_PerCellbyCondition.png |
| Annotation | annotation/annotation_m_umap_combined.png | 3_Ann_pt2/M/UMAP_annot_combined.png |
| Annotation | annotation/annotation_m_marker_dotplot.png | 3_Ann_pt2/M/Multiome_DotPlot_Markers.png |
| Annotation | annotation/annotation_m_cells_by_condition.png | 3_Ann_pt2/M/BarPlot_PerCellbyCondition.png |
| GSEA | gsea/gsea_f_myonuclei_cac_vs_ctrl.png | 8_GSEA_New/F/MyoOverall_CACvsCTRL_WithRibo/dotplots/GSEA_dotplot_Myonuclei_CAC_vs_CTRL.png |
| GSEA | gsea/gsea_f_myonuclei_cac_vs_pre.png | 8_GSEA_New/F/MyoOverall_CACvsPRE_WithRibo/dotplots/GSEA_dotplot_Myonuclei_CAC_vs_PRE.png |
| GSEA | gsea/gsea_f_myonuclei_pre_vs_ctrl.png | 8_GSEA_New/F/MyoOverall_PREvsCTRL_WithRibo/dotplots/GSEA_dotplot_Myonuclei_PRE_vs_CTRL.png |
| GSEA | gsea/gsea_m_myonuclei_cac_vs_ctrl.png | 8_GSEA/M/MyoOverall_CACvsCTRL_WithRibo/dotplots/GSEA_dotplot_Myonuclei_CAC_vs_CTRL.png |
| GSEA | gsea/gsea_m_myonuclei_cac_vs_pre.png | 8_GSEA/M/MyoOverall_CACvsPRE_WithRibo/dotplots/GSEA_dotplot_Myonuclei_CAC_vs_PRE.png |
| GSEA | gsea/gsea_m_myonuclei_pre_vs_ctrl.png | 8_GSEA/M/MyoOverall_PREvsCTRL_WithRibo/dotplots/GSEA_dotplot_Myonuclei_PRE_vs_CTRL.png |
| DEVIL GSEA | devil/devil_gsea_f_myonuclei_cac_vs_ctrl.png | 8_GSEA_devil_New/F/MyoOverall_CACvsCTRL_WithRibo/dotplots/GSEA_dotplot_Myonuclei_CAC_vs_CTRL.png |
| DEVIL GSEA | devil/devil_gsea_m_myonuclei_cac_vs_ctrl.png | 8_GSEA_devil_New/M/MyoOverall_CACvsCTRL_WithRibo/dotplots/GSEA_dotplot_Myonuclei_CAC_vs_CTRL.png |
| CCC / Scriabin | ccc/ccc_myoIIx_to_tcells_clusters.png | EXTRA/11_CCC_Scriabin/CCIM_CAC_Myo_IIx_to_TCells/CCIM_UMAP_by_clusters.png |
| CCC / Scriabin | ccc/ccc_myoIIx_to_tcells_sender.png | EXTRA/11_CCC_Scriabin/CCIM_CAC_Myo_IIx_to_TCells/CCIM_UMAP_by_sender_clusters.png |
| CCC / Scriabin | ccc/ccc_myoIIx_to_tcells_receiver.png | EXTRA/11_CCC_Scriabin/CCIM_CAC_Myo_IIx_to_TCells/CCIM_UMAP_by_receiver_clusters.png |
| CCC / Scriabin | ccc/ccc_myoIIx_to_tcells_lr_dotplot.png | EXTRA/11_CCC_Scriabin/CCIM_CAC_Myo_IIx_to_TCells_markers/DotPlot_topLR_cluster0.png |
Repository Map
| Top_level |
|---|
| 0_tbi_file.R |
| 0_tbi_file_NewData.R |
| 1_QualityControl.R |
| 1_QualityControl_NewData.R |
| 2_RemoveDoublets_Fem.R |
| 2_RemoveDoublets_Fem_NewData.R |
| 2_RemoveDoublets_Man.R |
| 2_RemoveDoublets_Man_NewData.R |
| 3_QC_ribosom.R |
| hdf5-1.14.3.tar.gz |
| NuovaPipeNo120 |
| Output |
| Poster |
| quarto_interactive |
All R scripts currently tracked in the analysis folders
| File |
|---|
| 0_tbi_file.R |
| 0_tbi_file_NewData.R |
| 1_QualityControl.R |
| 1_QualityControl_NewData.R |
| 2_RemoveDoublets_Fem.R |
| 2_RemoveDoublets_Fem_NewData.R |
| 2_RemoveDoublets_Man.R |
| 2_RemoveDoublets_Man_NewData.R |
| 3_QC_ribosom.R |
| NuovaPipeNo120/0_PseudoBulkxClusterF.R |
| NuovaPipeNo120/0_PseudoBulkxClusterM.R |
| NuovaPipeNo120/1_IntFem_NewData.R |
| NuovaPipeNo120/1_IntMan_NewData.R |
| NuovaPipeNo120/10_CCC_120_pt1.R |
| NuovaPipeNo120/10_CCC_120_pt2.R |
| NuovaPipeNo120/10_CCC_F.R |
| NuovaPipeNo120/10_CCC_F_perPaz.R |
| NuovaPipeNo120/10_CCC_M.R |
| NuovaPipeNo120/10_CCC_M_perPaz.R |
| NuovaPipeNo120/2_AnnFem_New.R |
| NuovaPipeNo120/2_AnnMan_New.R |
| NuovaPipeNo120/3_AnnFem_pt2_New.R |
| NuovaPipeNo120/3_AnnMan_pt2_New.R |
| NuovaPipeNo120/4_FiltGenes_F_NewData.R |
| NuovaPipeNo120/4_FiltGenes_M_NewData.R |
| NuovaPipeNo120/5_DEG_F_New.R |
| NuovaPipeNo120/5_DEG_M_New.R |
| NuovaPipeNo120/5_DEG_New.R |
| NuovaPipeNo120/6_DEA_devil_female.R |
| NuovaPipeNo120/6_DEA_devil_male.R |
| NuovaPipeNo120/7_GSEA_F.R |
| NuovaPipeNo120/7_GSEA_M_New.R |
| NuovaPipeNo120/8_GSEA_F_Dev.R |
| NuovaPipeNo120/8_GSEA_M_Dev.R |
| NuovaPipeNo120/8_Ribosomal_PCA_Heatmaps_MF.R |
| NuovaPipeNo120/ConfrontoLabelMan.R |
| NuovaPipeNo120/Dotplot.R |
| NuovaPipeNo120/Dotplot_F.R |
| NuovaPipeNo120/Dotplot_M.R |
| NuovaPipeNo120/Heatmap_F.R |
| NuovaPipeNo120/Heatmap_M.R |
| NuovaPipeNo120/HeatRibo.R |
| NuovaPipeNo120/old/0_bulkPCA_All.R |
| NuovaPipeNo120/old/0_bulkPCA_All_No120.R |
| NuovaPipeNo120/old/0_bulkPCA_F.R |
| NuovaPipeNo120/old/0_bulkPCA_F_NewData_NoC063.R |
| NuovaPipeNo120/old/0_bulkPCA_F_No120.R |
| NuovaPipeNo120/old/0_bulkPCA_M.R |
| NuovaPipeNo120/old/1_IntFem.R |
| NuovaPipeNo120/old/1_IntMan.R |
| NuovaPipeNo120/old/2_AnnFem.R |
| NuovaPipeNo120/old/2_AnnMan.R |
| NuovaPipeNo120/old/3_AnnFem_pt2.R |
| NuovaPipeNo120/old/3_AnnMan_pt2.R |
| NuovaPipeNo120/old/4_FiltGenes_F.R |
| NuovaPipeNo120/old/4_FiltGenes_M.R |
| NuovaPipeNo120/old/5_DEG_F.R |
| NuovaPipeNo120/old/5_DEG_M.R |
| NuovaPipeNo120/old/5_DEGs_M_skeletal.R |
| NuovaPipeNo120/old/5_DEGs_M_Undet.R |
| NuovaPipeNo120/old/9_GSEA_M.R |
| NuovaPipeNo120/ribo/EXTRA_Ribo_Dotplot_F.R |
| NuovaPipeNo120/ribo/EXTRA_Ribo_Dotplot_M.R |
| NuovaPipeNo120/ribo/EXTRA_Ribo_PCA_F.R |
| NuovaPipeNo120/ribo/EXTRA_Ribo_PCA_M.R |
| NuovaPipeNo120/scriabin/11_Scriabin_M_CAC_vs_CTRL_pt1.R |
| NuovaPipeNo120/scriabin/11_Scriabin_M_CAC_vs_CTRL_pt2.R |
| NuovaPipeNo120/scriabin/11_Scriabin_M_CAC_vs_CTRL_pt2_Global.R |
| NuovaPipeNo120/scriabin/11_Scriabin_M_CAC_vs_CTRL_pt2_NoGrafo.R |
| NuovaPipeNo120/scriabin/11_Scriabin_M_CAC_vs_CTRL_pt3.R |
| NuovaPipeNo120/scriabin/11_Scriabin_M_CAC_vs_CTRL_pt3_Global.R |
| NuovaPipeNo120/scriabin/11_Scriabin_M_CAC_vs_CTRL_pt4.R |
| NuovaPipeNo120/scriabin/11_Scriabin_M_CAC_vs_CTRL_pt4_Global.R |
| NuovaPipeNo120/scriabin/11_Scriabin_M_CTRL_vs_CAC_pt2.R |
| NuovaPipeNo120/scriabin/11_Scriabin_M_FiltGenes_pt2.R |
| NuovaPipeNo120/scriabin/13_CCIM_CellChatStyle_MyoIIx_to_TCells.R |
| NuovaPipeNo120/Umap_F.R |
| NuovaPipeNo120/Umap_M.R |
| NuovaPipeNo120/Upset_M.R |
| Poster/Annotazioni/clust2.R |
| Poster/Annotazioni/clust3.R |
| Poster/Annotazioni/CreateClusterExp.R |
| Poster/Annotazioni/provaClust.R |
| Poster/Annotazioni/provaClust2.R |
| Poster/Annotazioni/Seurat.R |
| Poster/Annotazioni/sgd.R |
| Poster/confronto_Fem.R |
| Poster/Seurat/1_IntFem_seurat_20k-1k-10.R |
| Poster/Seurat/1_IntFem_seurat_20k-1k-20.R |
| Poster/Seurat/1_IntFem_seurat_20k-1k-30.R |
| Poster/Seurat/1_IntFem_seurat_20k-2k-10.R |
| Poster/Seurat/1_IntFem_seurat_20k-2k-20.R |
| Poster/Seurat/1_IntFem_seurat_20k-2k-30.R |
| Poster/Seurat/1_IntFem_seurat_20k-3k-10.R |
| Poster/Seurat/1_IntFem_seurat_20k-3k-20.R |
| Poster/Seurat/1_IntFem_seurat_20k-3k-30.R |
| Poster/Seurat/1_IntFem_seurat_50k-1k-10.R |
| Poster/Seurat/1_IntFem_seurat_50k-1k-20.R |
| Poster/Seurat/1_IntFem_seurat_50k-1k-30.R |
| Poster/Seurat/1_IntFem_seurat_50k-2k-10.R |
| Poster/Seurat/1_IntFem_seurat_50k-2k-20.R |
| Poster/Seurat/1_IntFem_seurat_50k-2k-30.R |
| Poster/Seurat/1_IntFem_seurat_50k-3k-10.R |
| Poster/Seurat/1_IntFem_seurat_50k-3k-20.R |
| Poster/Seurat/1_IntFem_seurat_50k-3k-30.R |
| Poster/Seurat/1_IntFem_seurat_Max-1k-10.R |
| Poster/Seurat/1_IntFem_seurat_Max-1k-20.R |
| Poster/Seurat/1_IntFem_seurat_Max-1k-30.R |
| Poster/Seurat/1_IntFem_seurat_Max-2k-10.R |
| Poster/Seurat/1_IntFem_seurat_Max-2k-20.R |
| Poster/Seurat/1_IntFem_seurat_Max-2k-30.R |
| Poster/Seurat/1_IntFem_seurat_Max-3k-10.R |
| Poster/Seurat/1_IntFem_seurat_Max-3k-20.R |
| Poster/Seurat/1_IntFem_seurat_Max-3k-30.R |
| Poster/sgdGMF/annotazione/0_Preprocessing_Integration_Fem.R |
| Poster/sgdGMF/annotazione/0_Preprocessing_Integration_M.R |
| Poster/sgdGMF/annotazione/2_IntFem_30.R |
| Poster/sgdGMF/annotazione/2_IntM_30.R |
| Poster/sgdGMF/annotazione/3_Ann_M_30.R |
| Poster/sgdGMF/annotazione/3_AnnFem30.R |
| Poster/sgdGMF/annotazione/4_AnnFem_pt2.R |
| Poster/sgdGMF/sgd_20k-1k-10.R |
| Poster/sgdGMF/sgd_20k-1k-20.R |
| Poster/sgdGMF/sgd_20k-1k-30.R |
| Poster/sgdGMF/sgd_20k-2k-10.R |
| Poster/sgdGMF/sgd_20k-2k-20.R |
| Poster/sgdGMF/sgd_20k-2k-30.R |
| Poster/sgdGMF/sgd_20k-3k-10.R |
| Poster/sgdGMF/sgd_20k-3k-20.R |
| Poster/sgdGMF/sgd_20k-3k-30.R |
| Poster/sgdGMF/sgd_50k-1k-10.R |
| Poster/sgdGMF/sgd_50k-1k-20.R |
| Poster/sgdGMF/sgd_50k-1k-30.R |
| Poster/sgdGMF/sgd_50k-2k-10.R |
| Poster/sgdGMF/sgd_50k-2k-20.R |
| Poster/sgdGMF/sgd_50k-2k-30.R |
| Poster/sgdGMF/sgd_50k-3k-10.R |
| Poster/sgdGMF/sgd_50k-3k-20.R |
| Poster/sgdGMF/sgd_50k-3k-30.R |
| Poster/sgdGMF/sgd_Max-1k-10.R |
| Poster/sgdGMF/sgd_Max-1k-20.R |
| Poster/sgdGMF/sgd_Max-1k-30.R |
| Poster/sgdGMF/sgd_Max-2k-10.R |
| Poster/sgdGMF/sgd_Max-2k-20.R |
| Poster/sgdGMF/sgd_Max-2k-30.R |
| Poster/sgdGMF/sgd_Max-3k-10.R |
| Poster/sgdGMF/sgd_Max-3k-20.R |
| Poster/sgdGMF/sgd_Max-3k-30.R |
Discussion Questions
QC
Which samples or clusters are most sensitive to the filtering choices?
Annotation
Which cell-state labels are strong enough for the main story, and which should remain provisional?
Sex-specificity
Which signals replicate in female and male analyses, and which are biologically plausible as sex-specific?
Mechanism
Which pathway or communication signal deserves validation first?
Meeting Notes
Use this section during live discussion.
- Main concern raised:
- Figure requested:
- Follow-up analysis:
- Decision: